Myeloids subset analysis
Last compiled on 17 November, 2022
We considered five major cell subsets: (1) myeloid, (2) T, (3) B and plasma, (4) stromal, and (5) epithelial cells. We classified cells into these subsets according to their gene expression. We consider in this file the cells classified as epithelium. We reanalyze and QC the data,and look for the refined clusters.
Load libraries
library(Seurat)
library(plyr)
library(dplyr)
library(ggplot2)
library(DropletUtils)
library(celda)
library(SingleCellExperiment)
library(scater)
library(scran)
library(scDblFinder)
library(viridis)
library(MASS)
library(patchwork)
library(readr)
library(clustree)Load extra functions and sources
get_density <- function(x, y, ...) { # function from https://slowkow.com/notes/ggplot2-color-by-density/
dens <- MASS::kde2d(x, y, ...)
ix <- findInterval(x, dens$x)
iy <- findInterval(y, dens$y)
ii <- cbind(ix, iy)
return(dens$z[ii])
}
fancy_scientific <- function(l) { # function from https://stackoverflow.com/a/24241954
# turn in to character string in scientific notation
l <- format(l, scientific = TRUE)
# quote the part before the exponent to keep all the digits
l <- gsub("^(.*)e", "'\\1'e", l)
# turn the 'e+' into plotmath format
l <- gsub("e", "%*%10^", l)
# return this as an expression
parse(text=l)
}
setwd("~/000_GitHub/ibd-bcn_single_cell")
source('source/functions_scrnaseq.R')
source('source/colors.R')Load the data
setwd('~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/')
myeloids <- readRDS('SUBSETS/FROM_SAMPLES_TOGETHER/myeloids.RDS')Reanalysis
These are the clusters we selected as myeloid cells in the “02_Data_to_subsets” file.
DimPlot(myeloids, label=T, group.by = 'RNA_snn_res.0.1')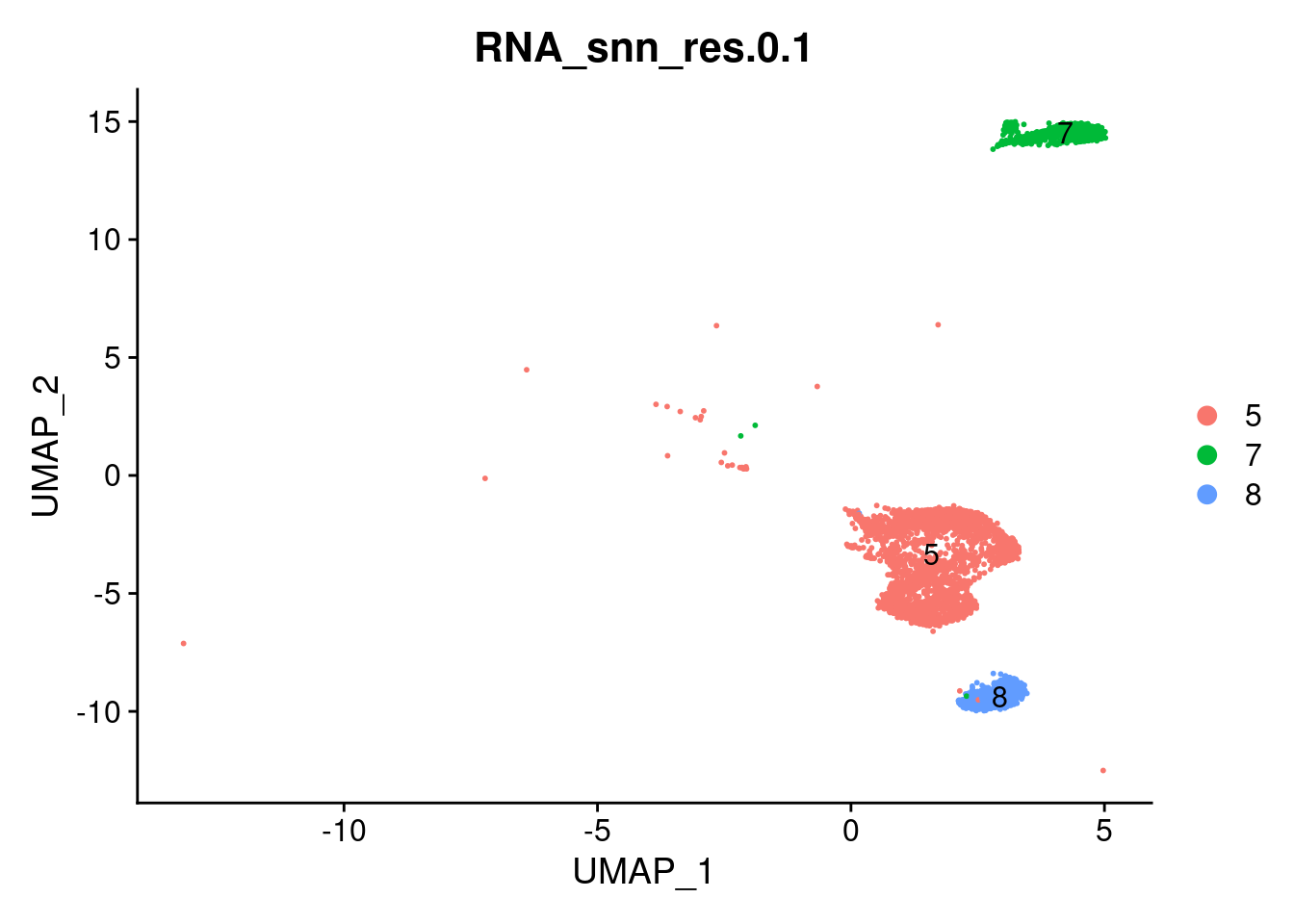
QC
MT-gene expression
Distribution of cells in scatter plot (nfeatures/percent.mt) to check where are the majority of our cells. We can see the dots colored by density using the function get_density shown in Load extra functions and sources.
meta <- myeloids@meta.data
meta$density <- get_density(meta$percent.mt, meta$nFeature_RNA, n = 100)
ggplot(meta) +
geom_point(aes(percent.mt, nFeature_RNA, color = density), size = 0.2) +
scale_color_viridis() + theme_classic() +
scale_y_log10()+
geom_vline(xintercept = 25, linetype = 2, color = 'gray')+
theme(text = element_text( size = 12),
axis.title = element_text( size = 12),
legend.text = element_blank())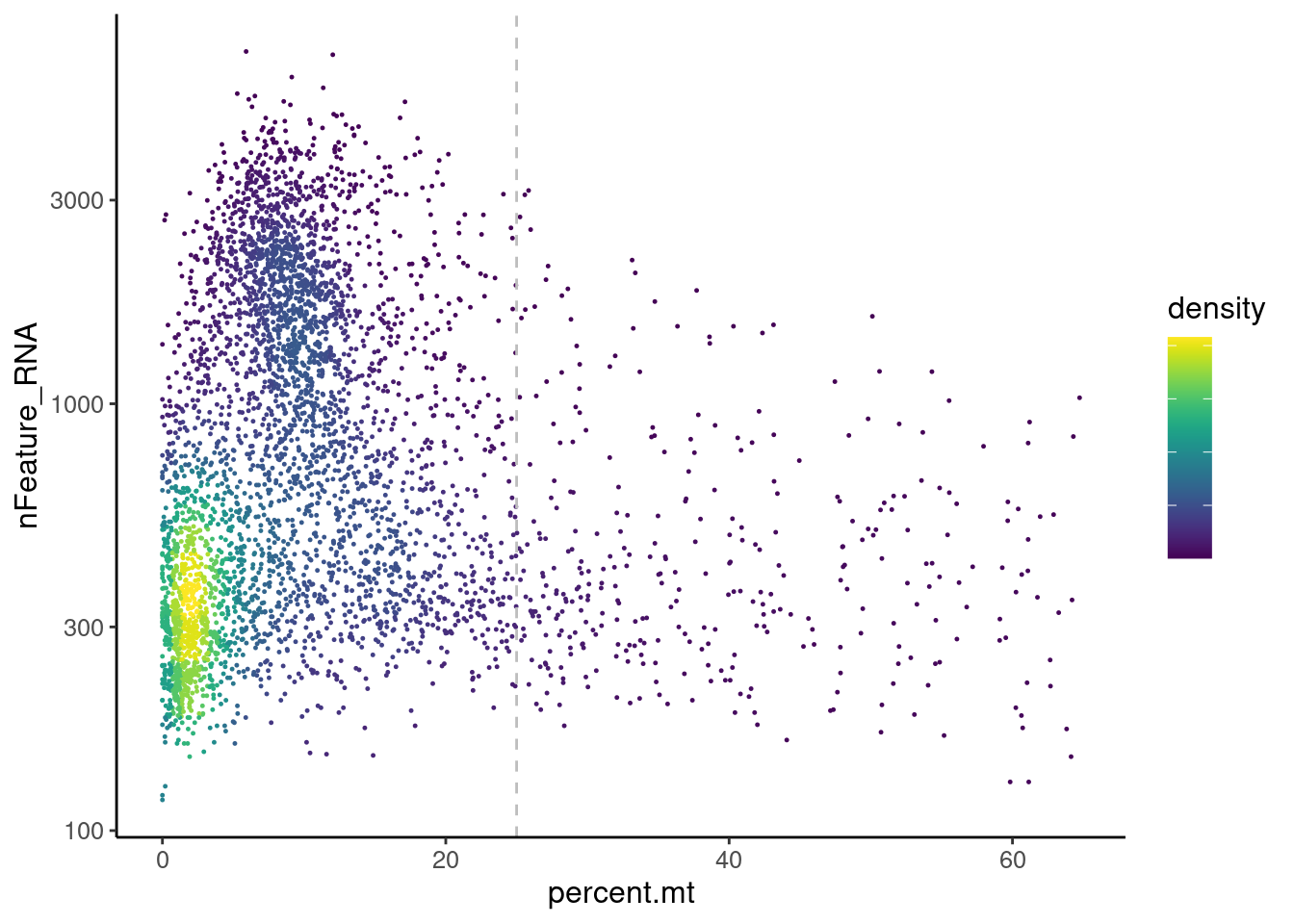
We analyzed the data using different cutoffs for the percent.mt (data not shown) and finally decided that 25% was a reasonable choice for this dataset.
myeloids <- myeloids[,myeloids$percent.mt < 25]Non-myeloid genes
Right now myeloids has 28287 features across 3964 cells.
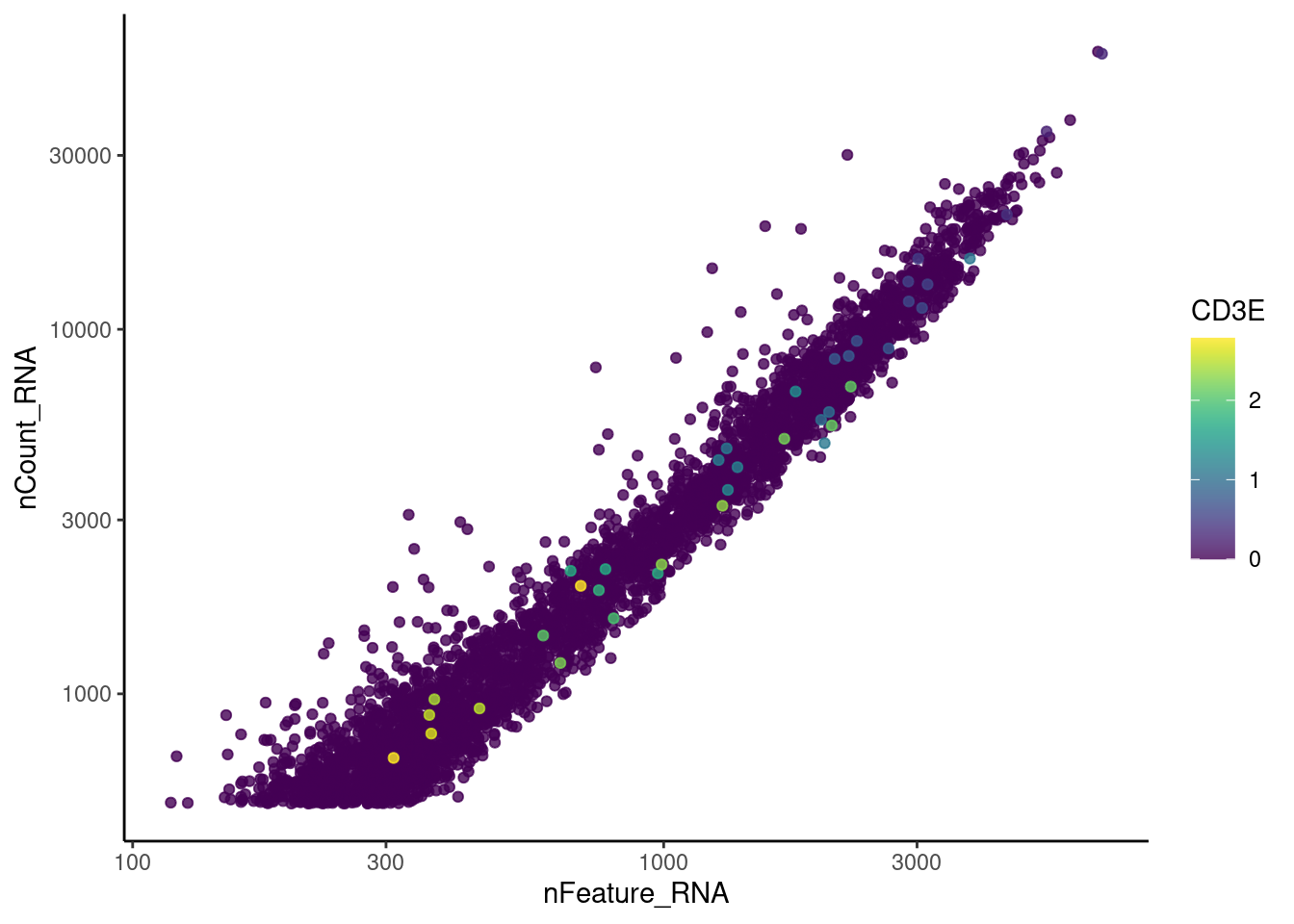
Nevertheless, many express genes that we know should not be expressed by
myeloid cells. We will remove those cells.
genes <- c('CD3E','CD3D','CD3G', 'THY1', 'DERL3', 'MS4A1')
for (gene in genes) {
jd <- FetchData(myeloids, vars = c('nCount_RNA', 'nFeature_RNA', gene))
jd <- jd[order(jd[,ncol(jd)]),]
k <- ggplot(jd, mapping = aes_string(x = 'nFeature_RNA', y = 'nCount_RNA', color = gene))+
geom_point() + theme_classic() + scale_color_viridis(alpha = 0.8) + scale_y_log10() + scale_x_log10()
cat("#### ", gene, "\n"); print(k); cat("\n\n")
}CD3E
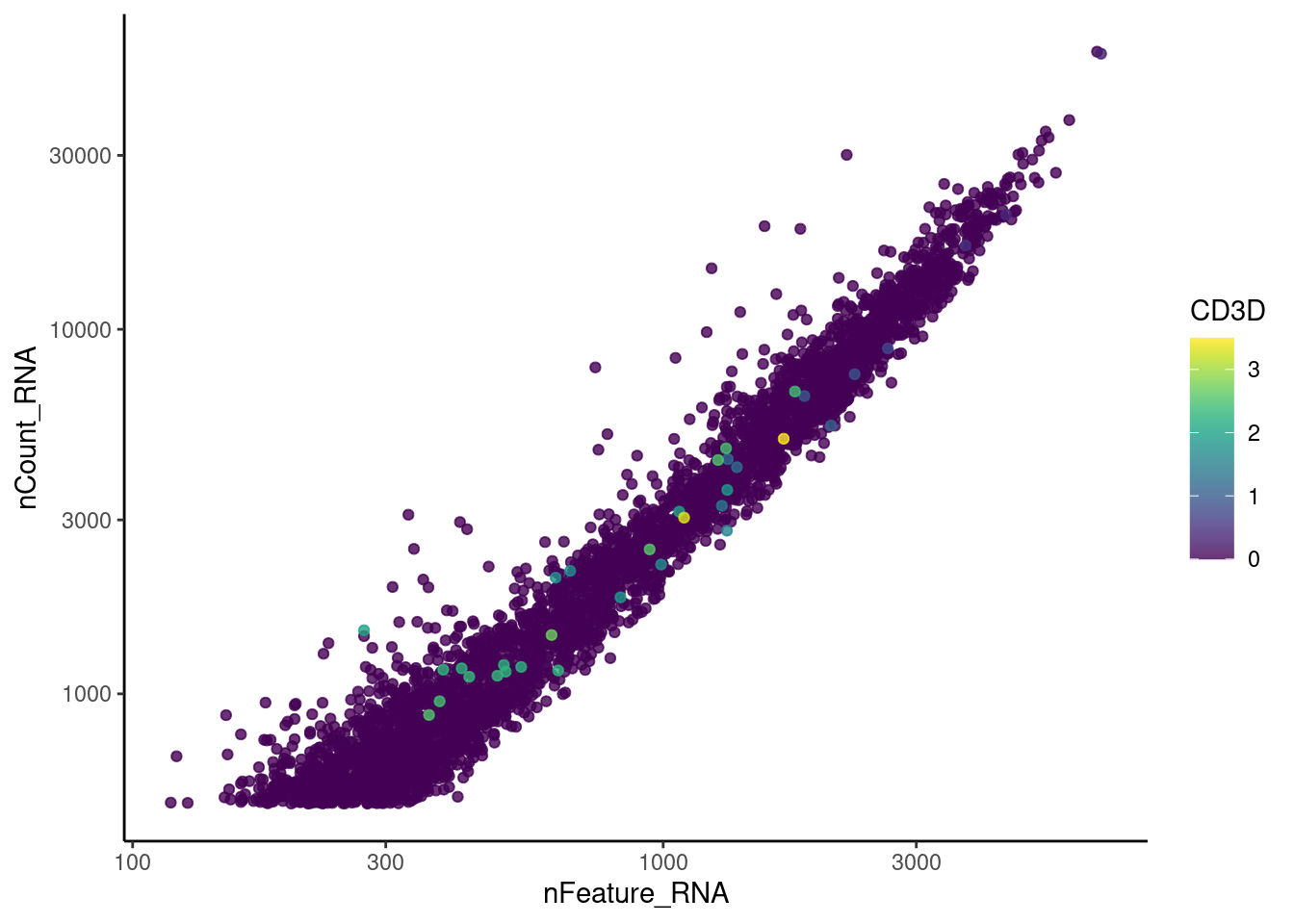
CD3D
CD3G
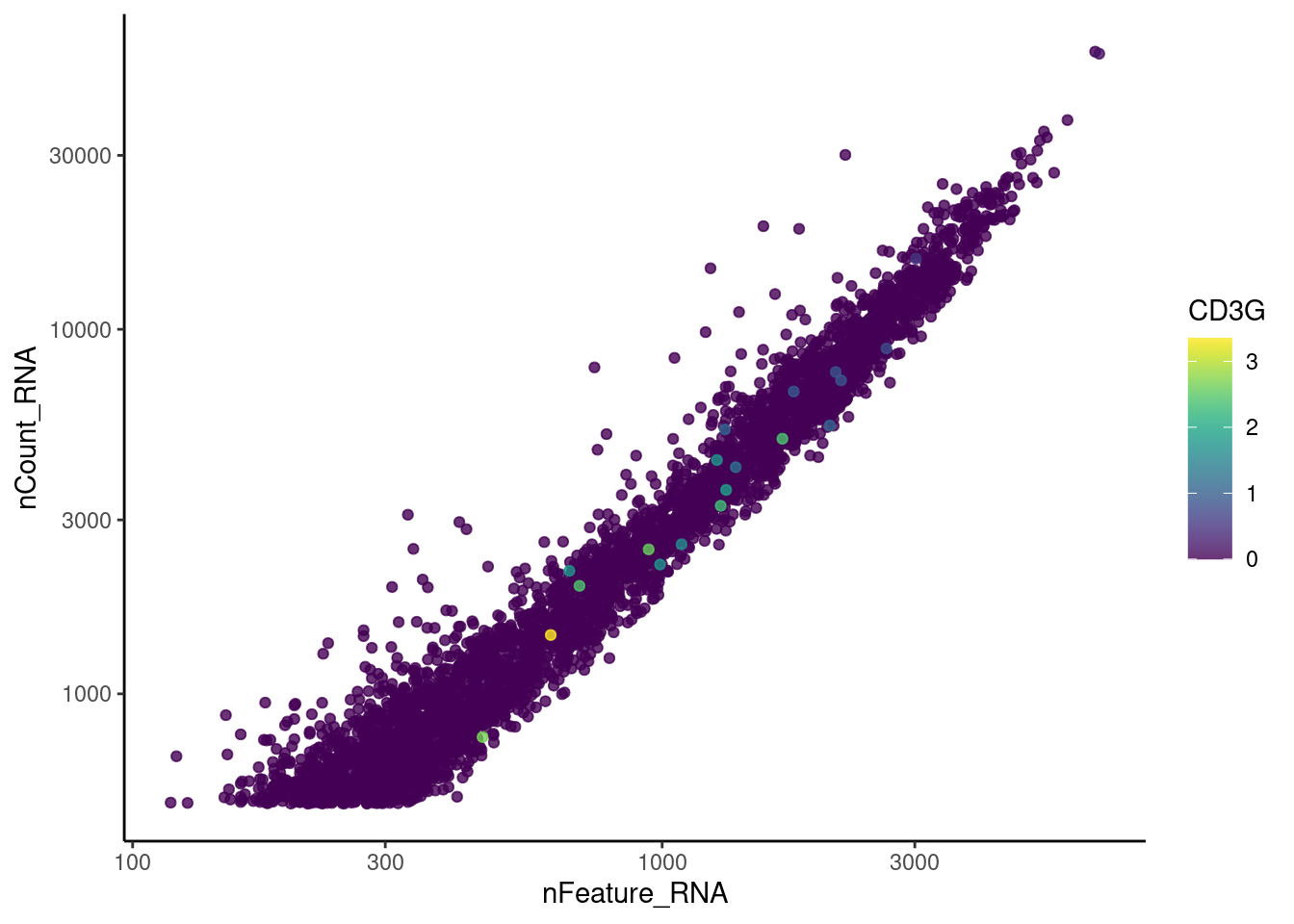
THY1
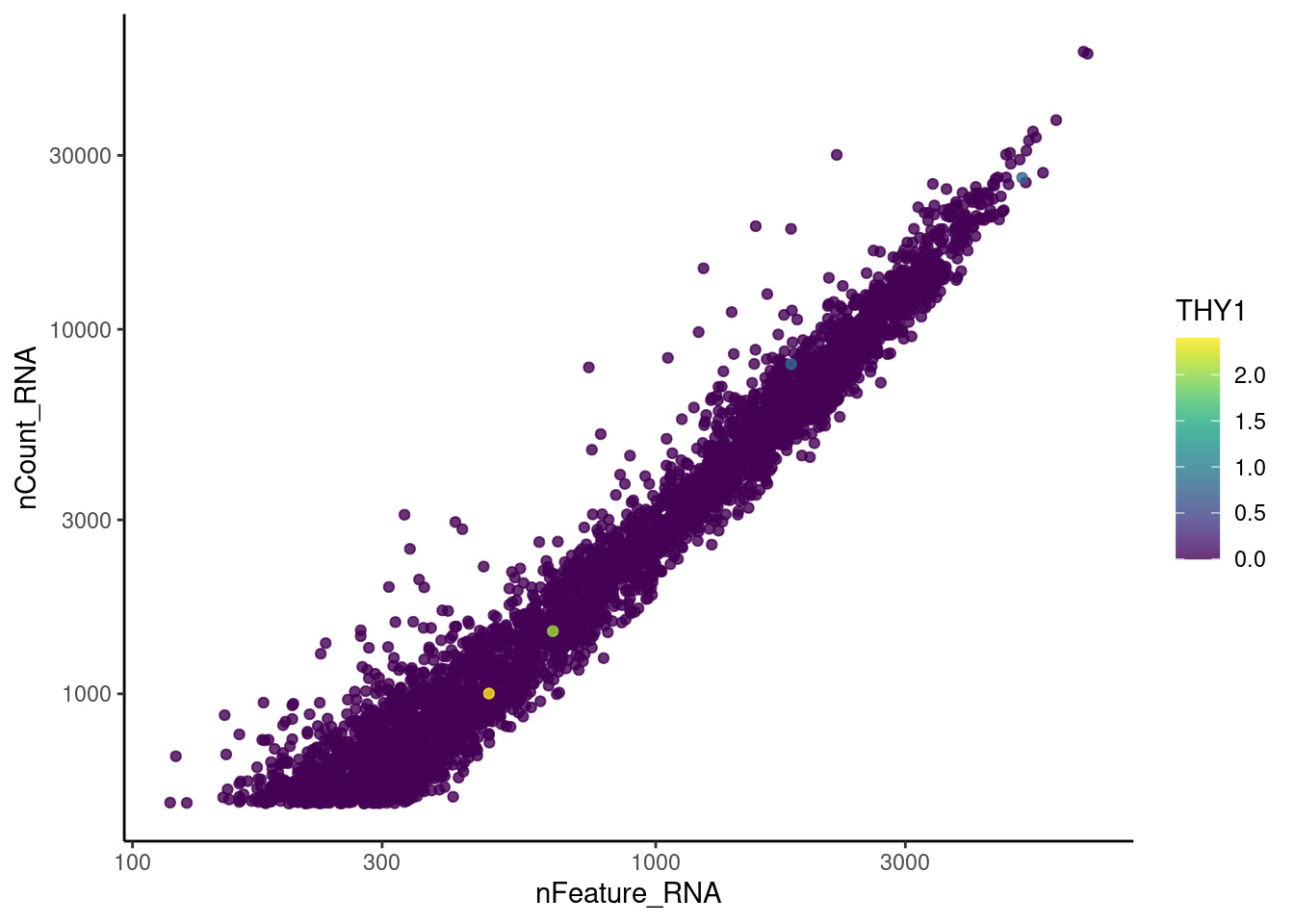
DERL3
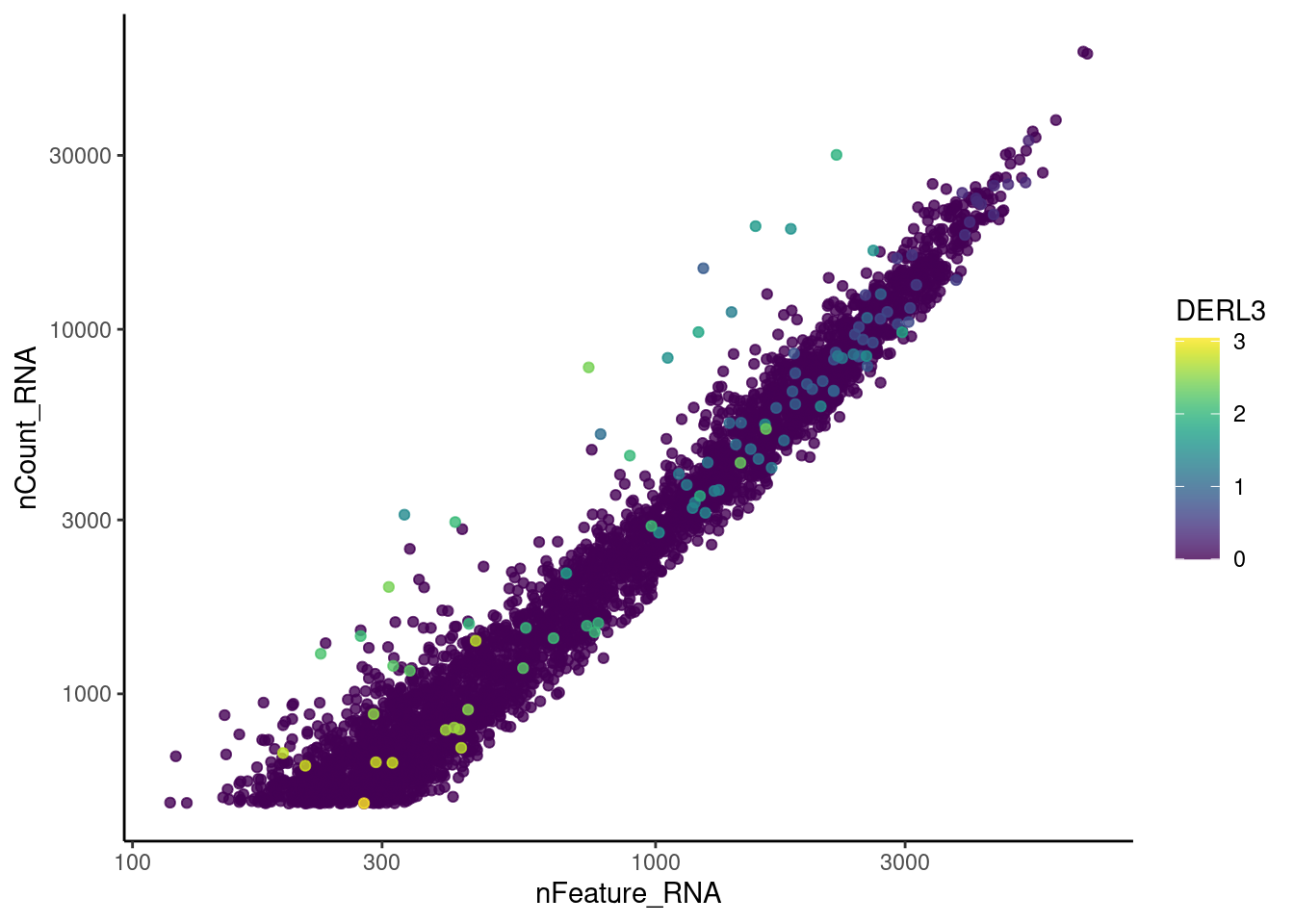
MS4A1
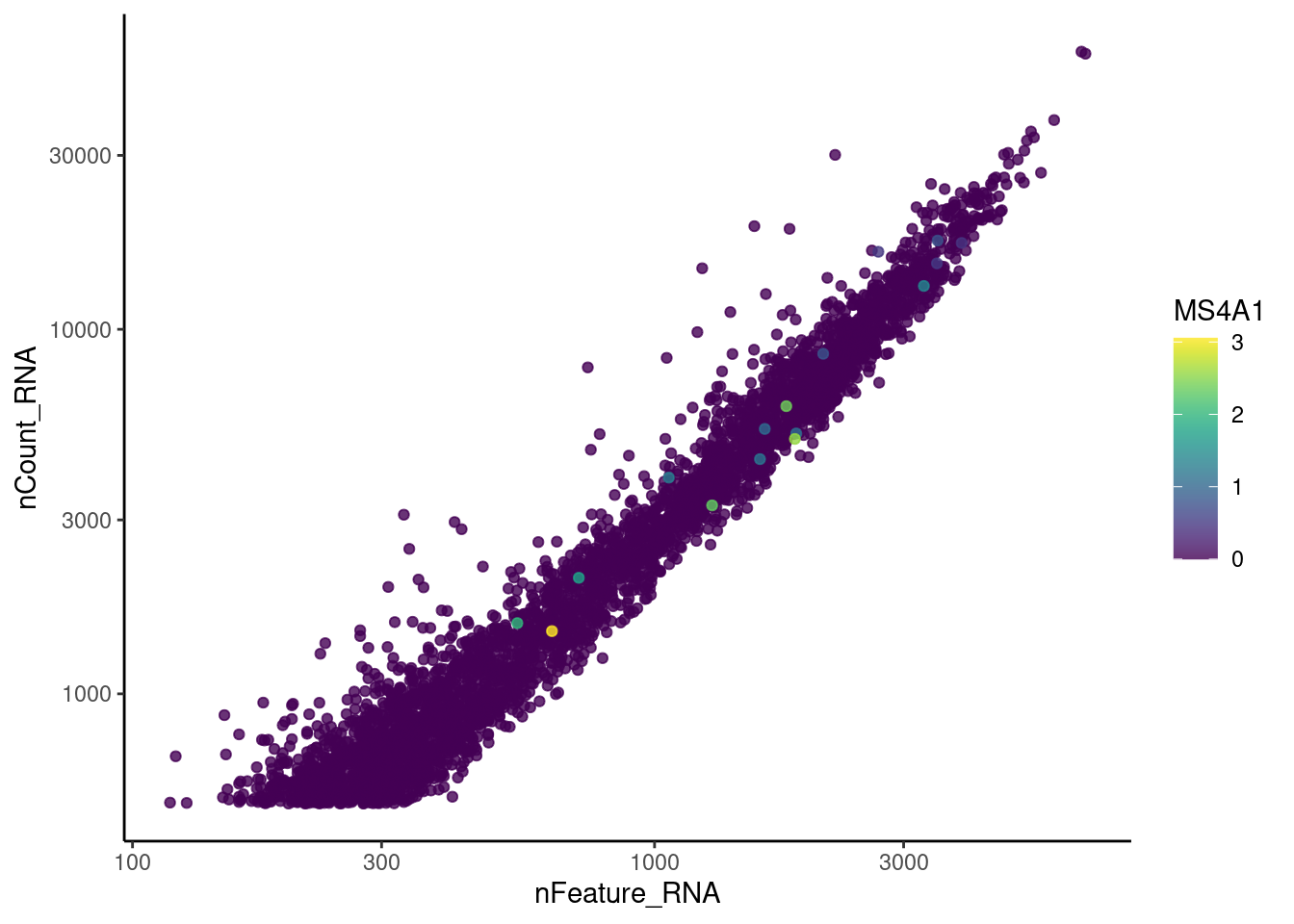
cat("#### code removal \n")code removal
counts <- myeloids@assays$RNA@counts
p <- grep("CD3E$|CD3D$|CD3G$|MS4A1$|DERL3|CD79A$|THY1$|EPCAM$",rownames(myeloids))
pp <- which(Matrix::colSums(counts[p,])>0)
xx <-setdiff(colnames(myeloids), names(pp))
myeloids <- subset(myeloids,cells = xx)
cat('\n\n')Ig genes signal
We have observed that due to normalization, there’s high Ig gene expression in cells that did not have high counts of Ig genes. As an example:
a <- FeaturePlot(myeloids, features = 'IGHA1', slot = 'counts') + labs(title='counts') + theme_classic()
b <- FeaturePlot(myeloids, features = 'IGHA1', slot = 'data') + labs(title= 'data') + theme_classic()
wrap_plots(a,b, nrow = 1)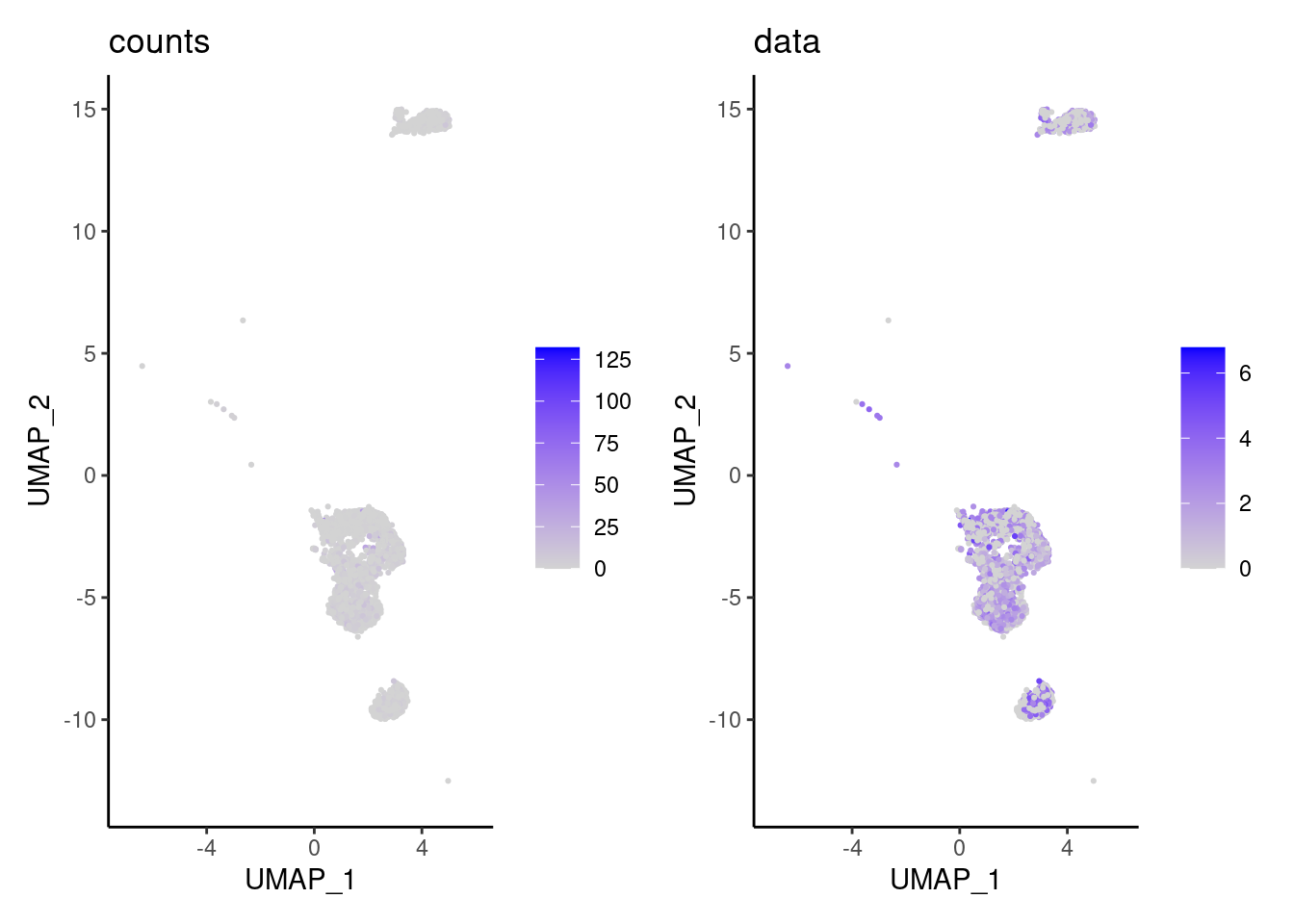
We remove the Ig genes from the dataset in all subsets except for the plasma and B cells subset.
gg <- rownames(myeloids)[c(grep("^IGH",rownames(myeloids)),
grep("^IGK", rownames(myeloids)),
grep("^IGL", rownames(myeloids)))]
genes <- setdiff(rownames(myeloids),gg)
myeloids <- subset(myeloids,features = genes)Genes without expression
We remove the genes without expression from the dataset.
counts <- myeloids@assays$RNA@counts
pp <- which(Matrix::rowSums(counts)==0)
xx <-setdiff(rownames(myeloids), names(pp))
myeloids <- subset(myeloids, features = xx)Dimension reduction
We tried different values of PC for the UMAP generation and Louvain clusterization. Finally, we considered 18 PCs for the analysis.
myeloids <- seurat_to_pca(myeloids)
a <- ElbowPlot(myeloids, ndims = 100) +
geom_vline(xintercept = 18, colour="#BB0000", linetype = 2)+
labs(title = paste0('Elbowplot')) + theme_classic()
myeloids <- FindNeighbors(myeloids, dims = 1:18)
myeloids <-RunUMAP(myeloids, dims=1:18)
b <- DimPlot(myeloids, group.by = 'sample_name') +
labs(title = '18 PCS') +
theme_classic() +
theme(legend.position = 'bottom')
(a+b) / guide_area() +
plot_layout(heights = c(0.7,0.3),guides = 'collect')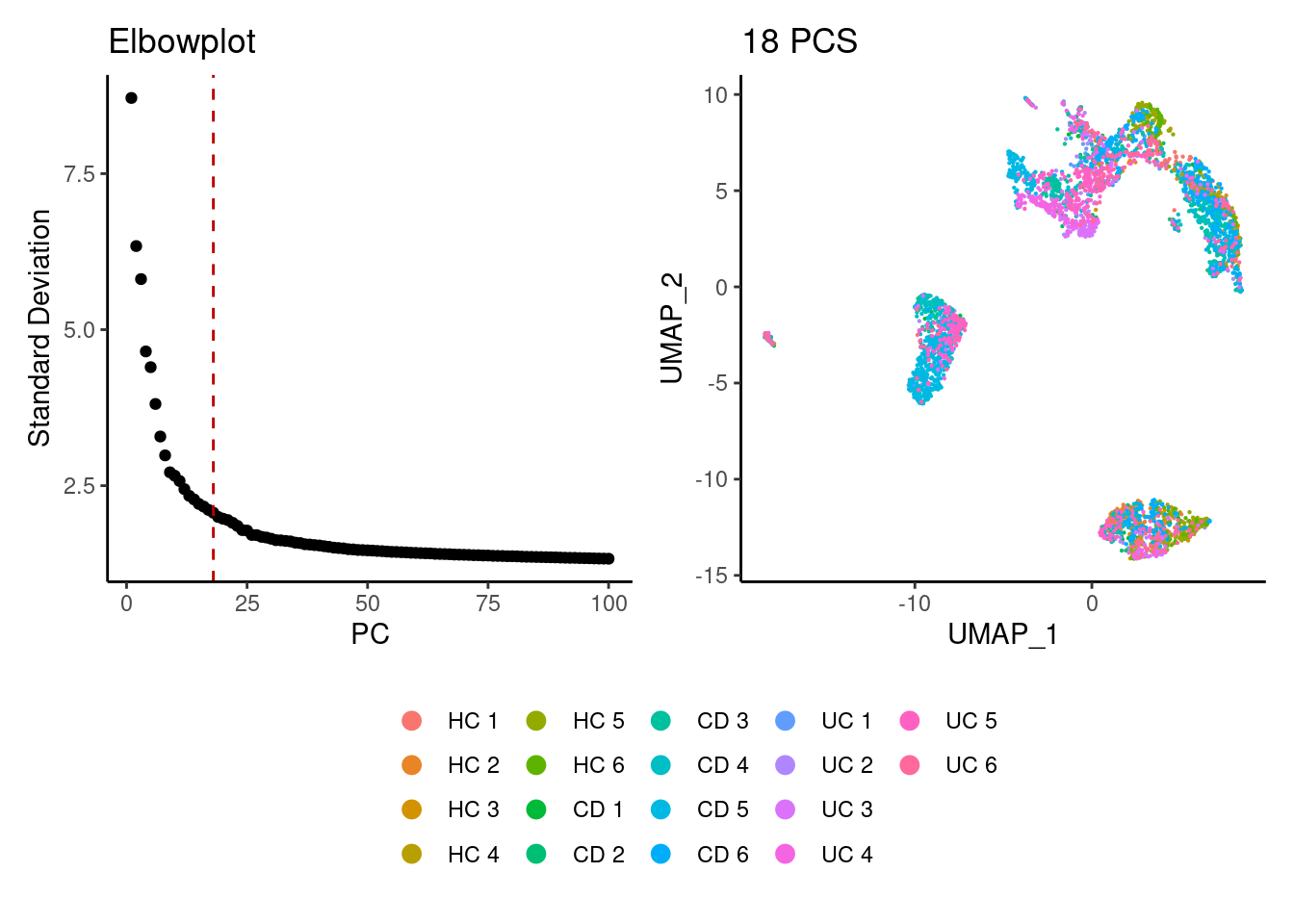
Louvain Clusterization
# dir.create('Markers')
# dir.create('Markers/Myeloids')
path <- '~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/SUBSETS/Markers'
setwd(path)
myeloids <- resolutions(myeloids,
workingdir = path,
title = 'Myeloids/Markers_Myeloids_')
path <- '~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/SUBSETS/ON_THEIR_OWN/'
setwd(path)
saveRDS(myeloids, file = paste0(path,'myeloids_filtered.RDS'))Dimplot Resolutions
path <- '~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/SUBSETS/ON_THEIR_OWN/'
myeloids <- readRDS(file = paste0(path,'myeloids_filtered.RDS'))
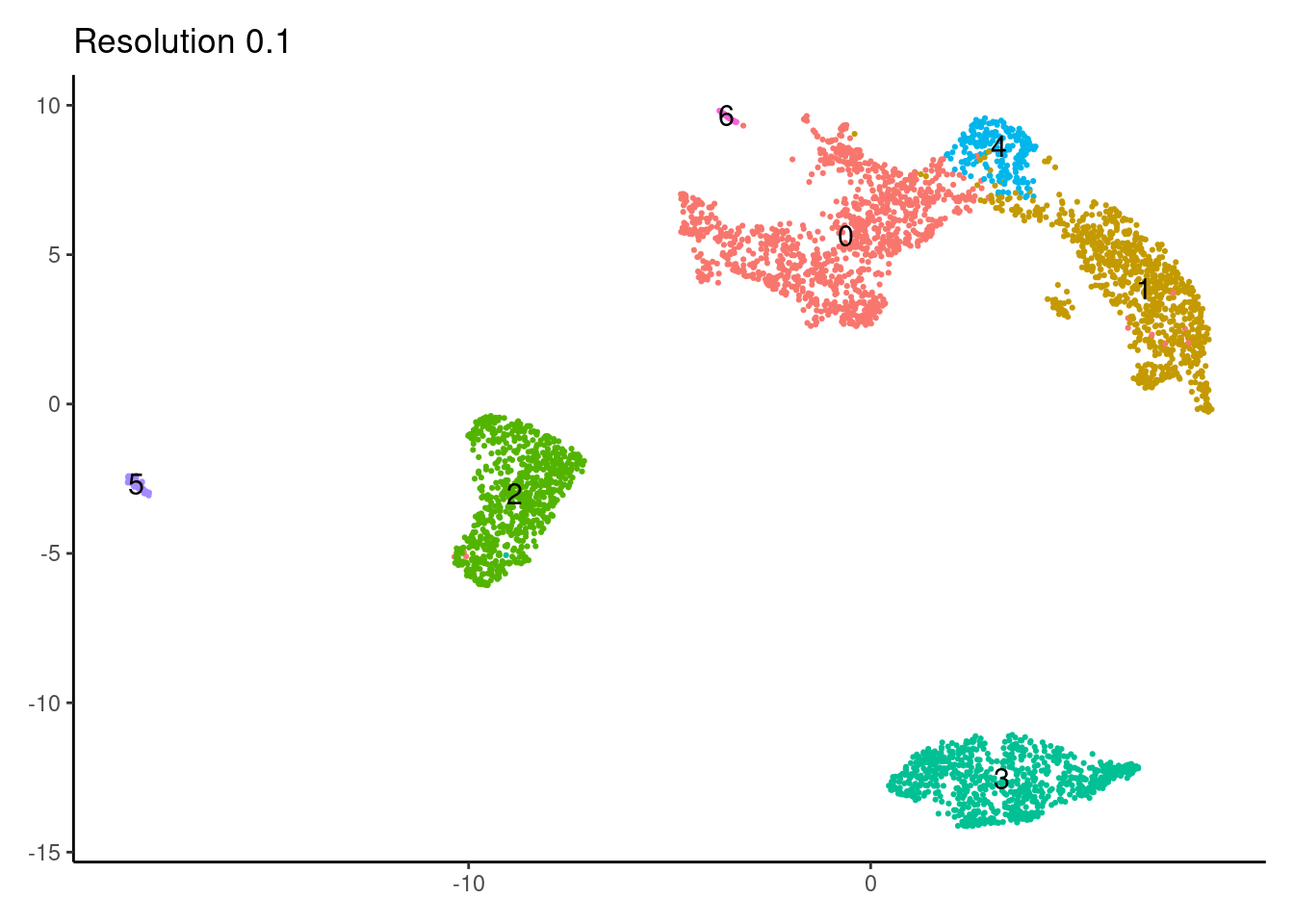
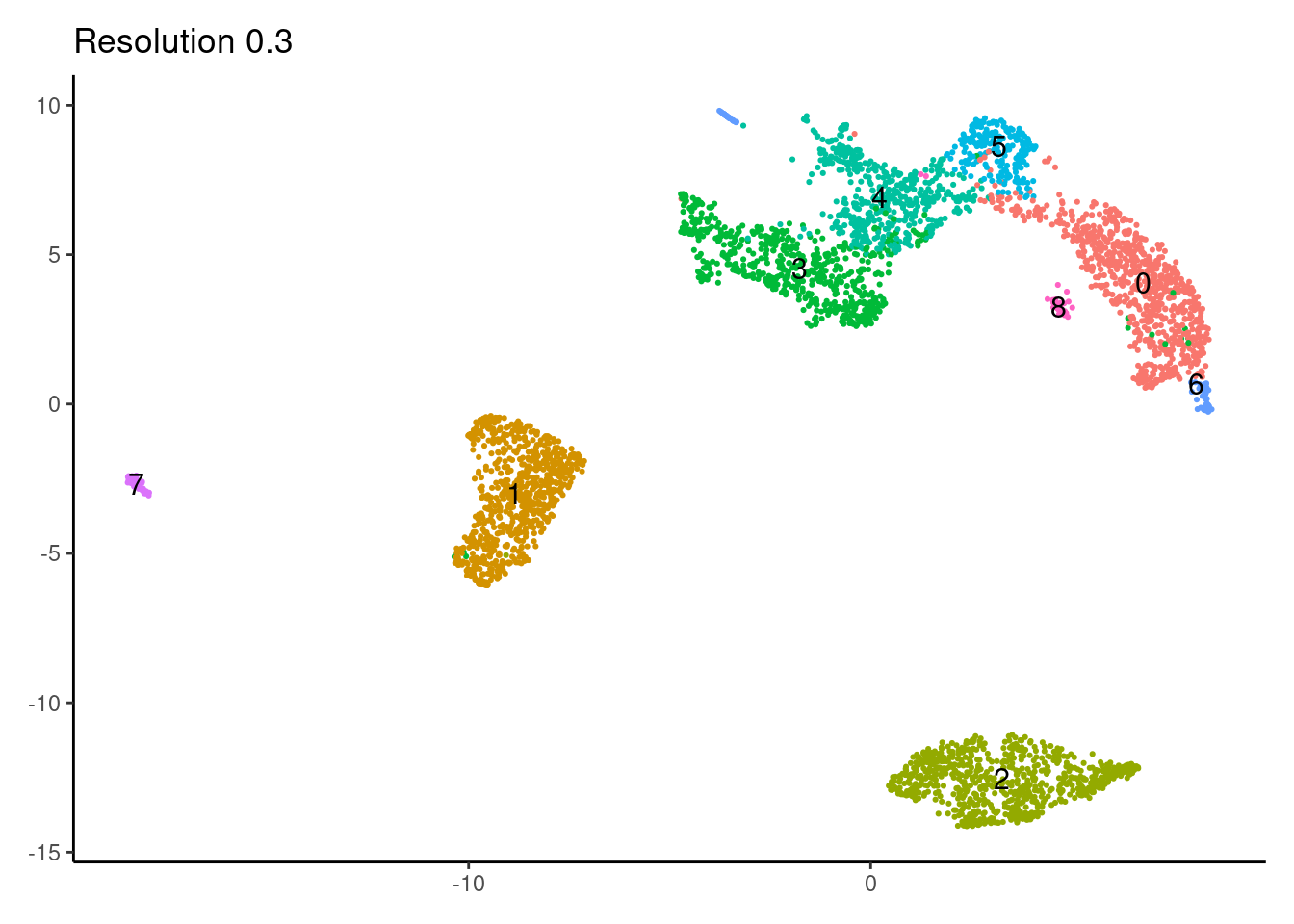
for(i in c('0.1','0.3','0.5','0.7','0.9','1.1','1.3','1.5')){
j <-DimPlot(myeloids, group.by = paste0('RNA_snn_res.',i), label=T) +
theme_classic() +
theme(legend.position = 'none', axis.title = element_blank()) +
labs(title = paste0('Resolution ',i))
cat("##### ", i, "\n"); print(j); cat("\n\n")
}0.1
0.3
0.5
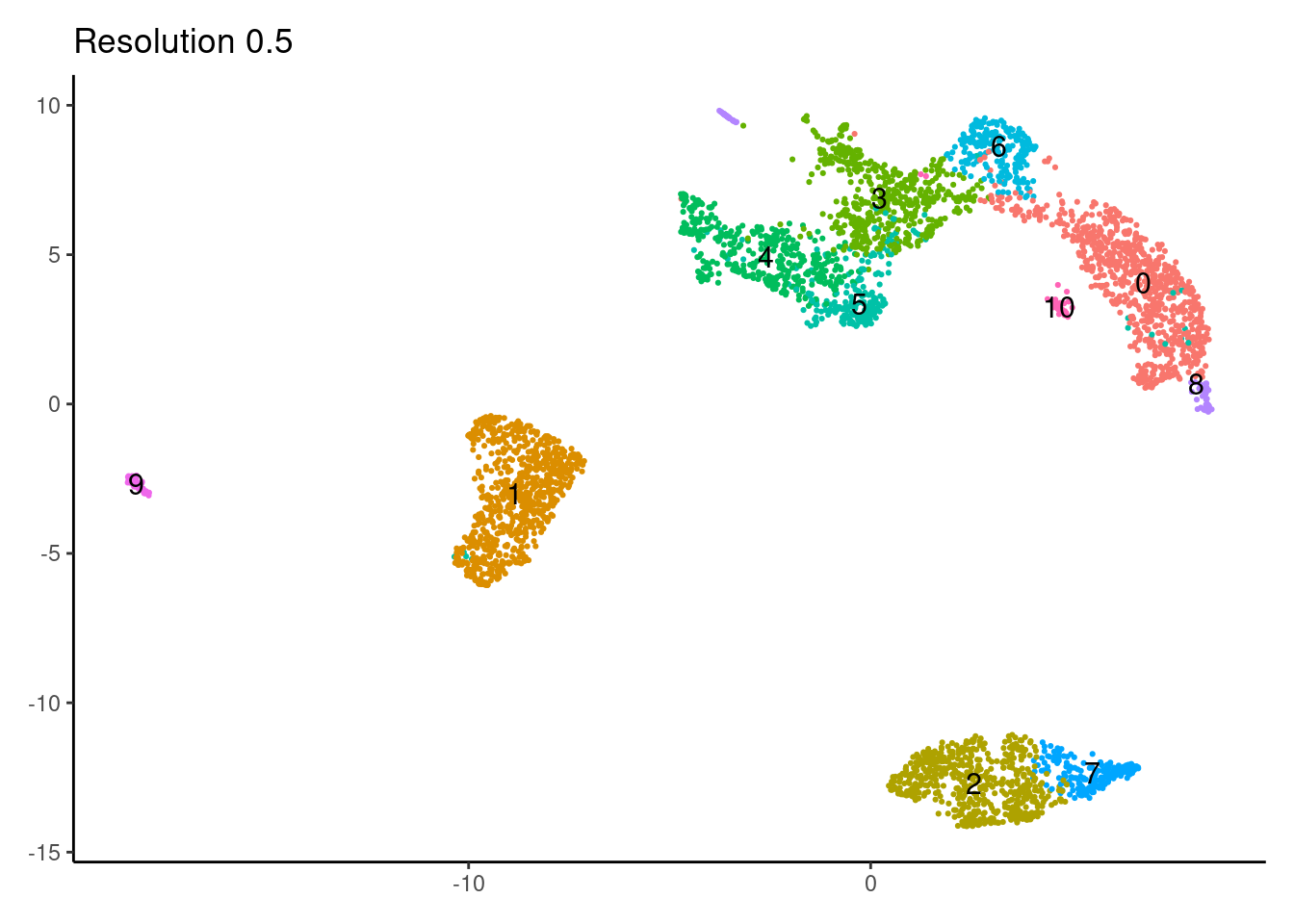
0.7
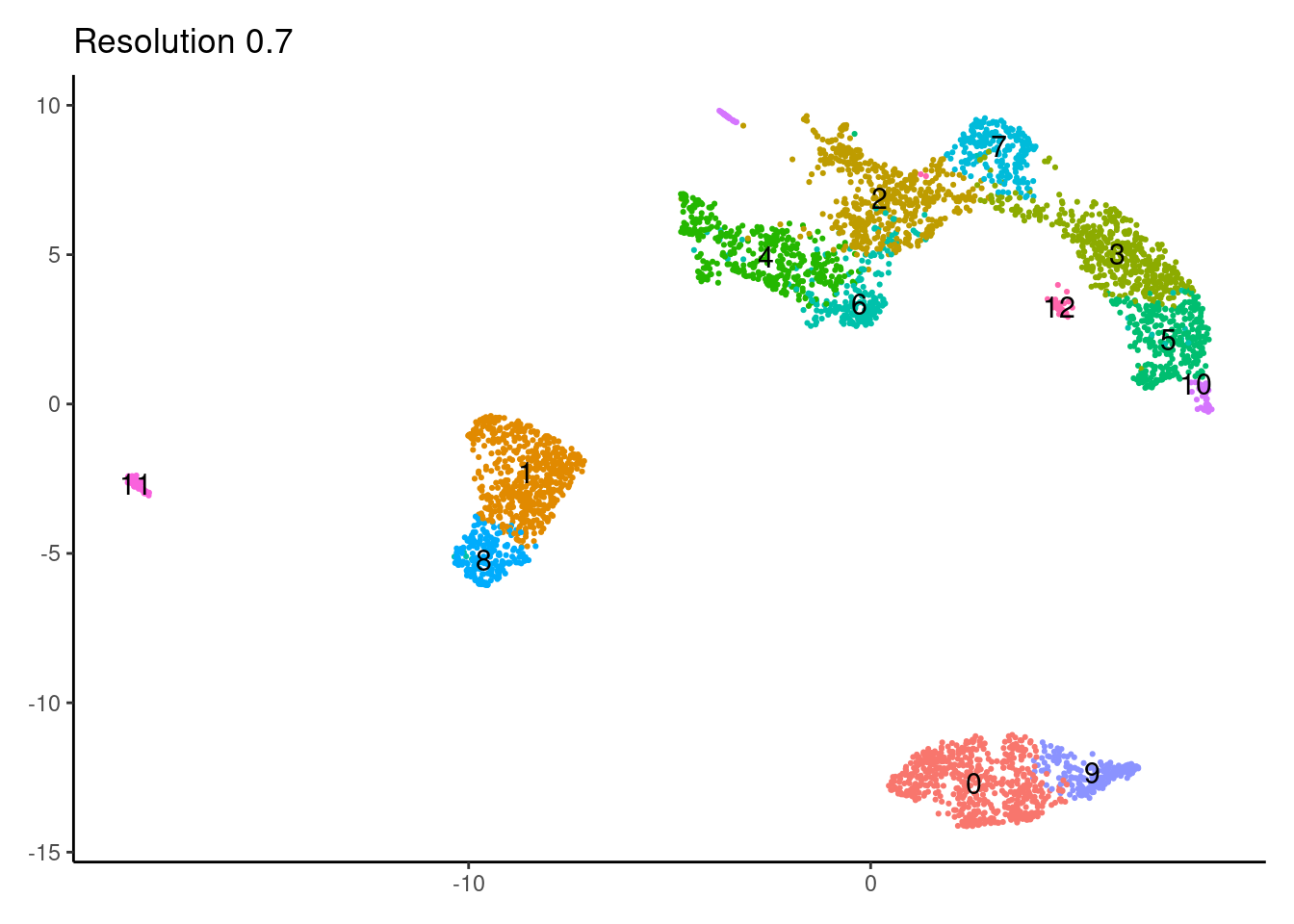
0.9
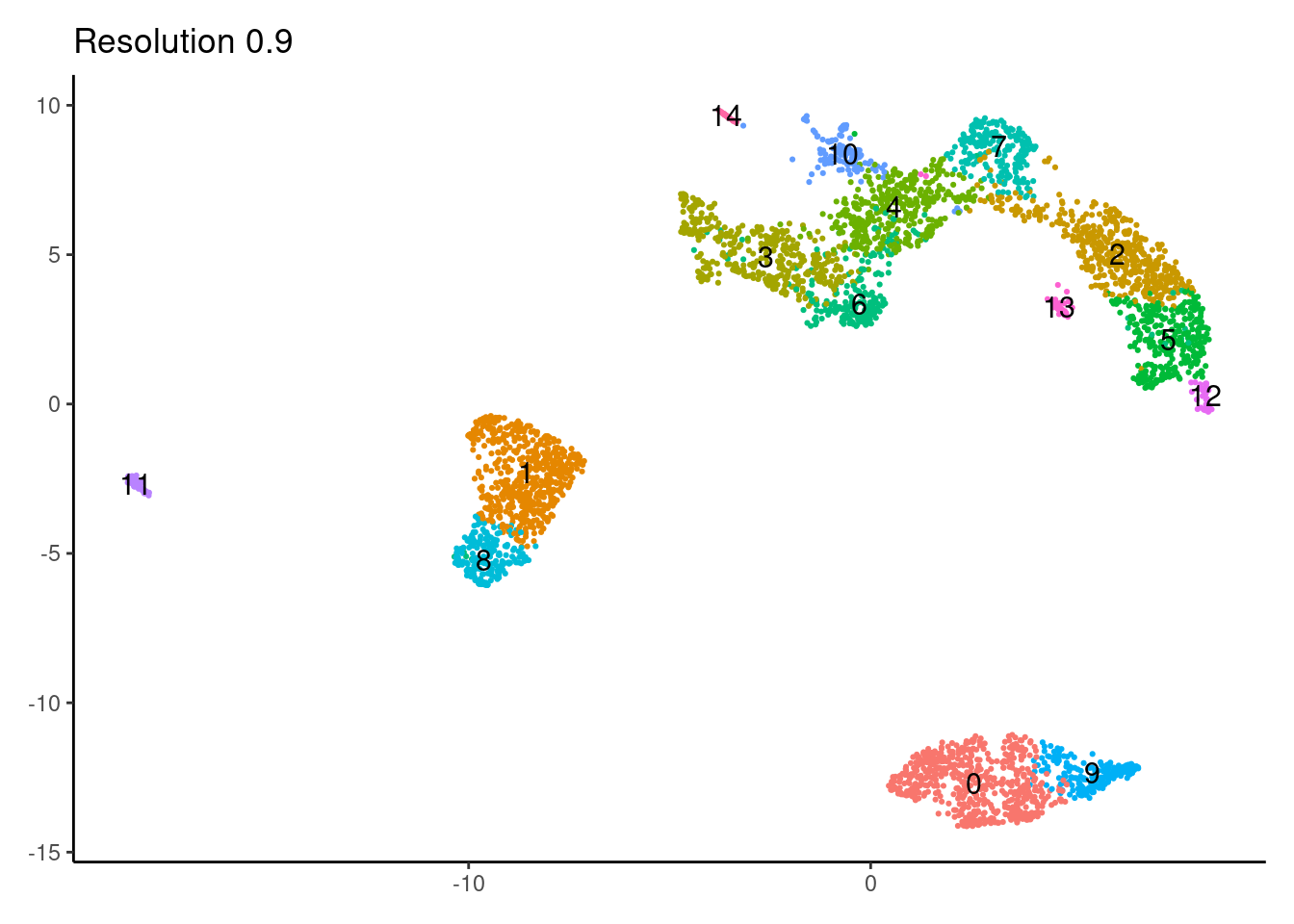
1.1
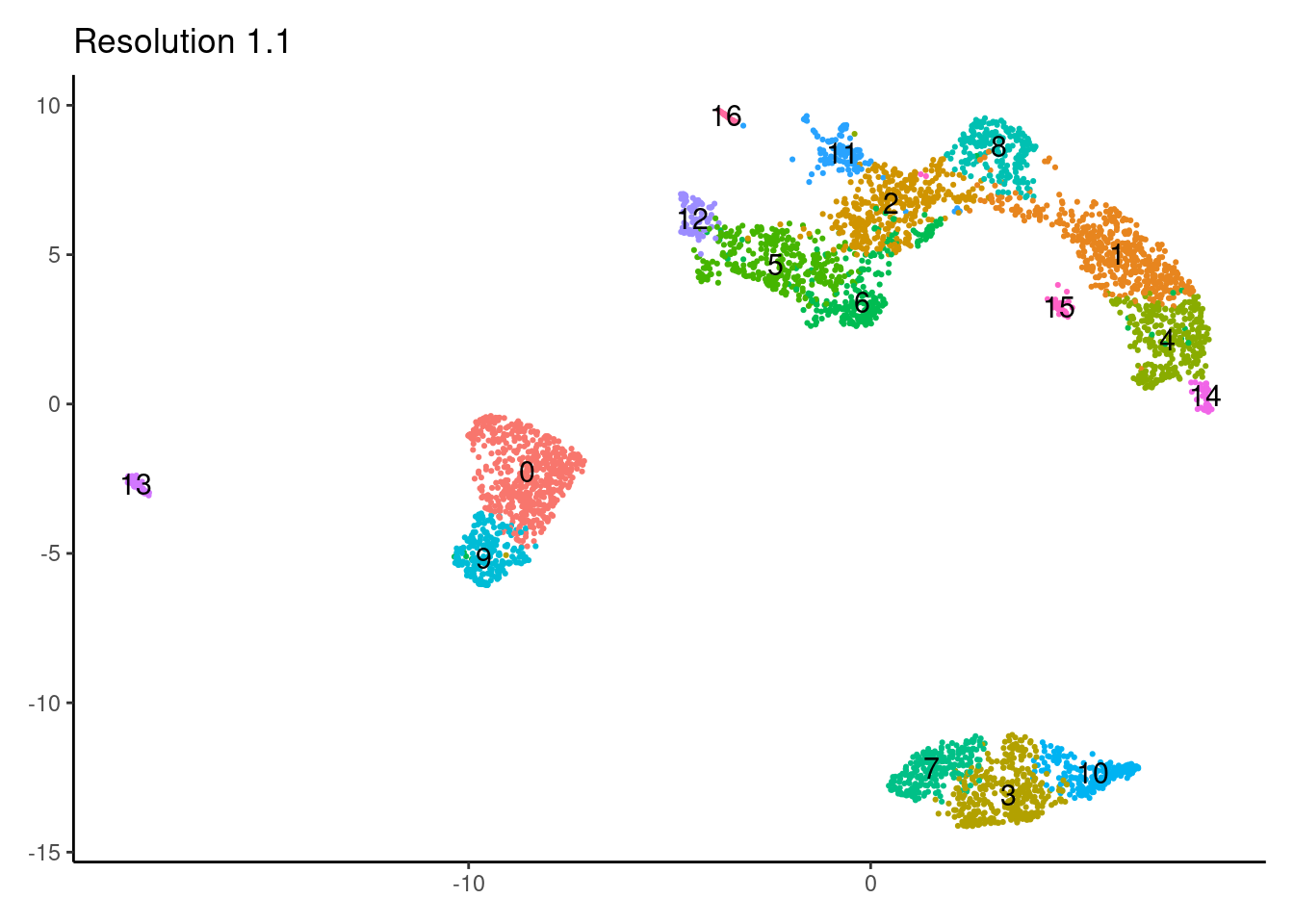
1.3
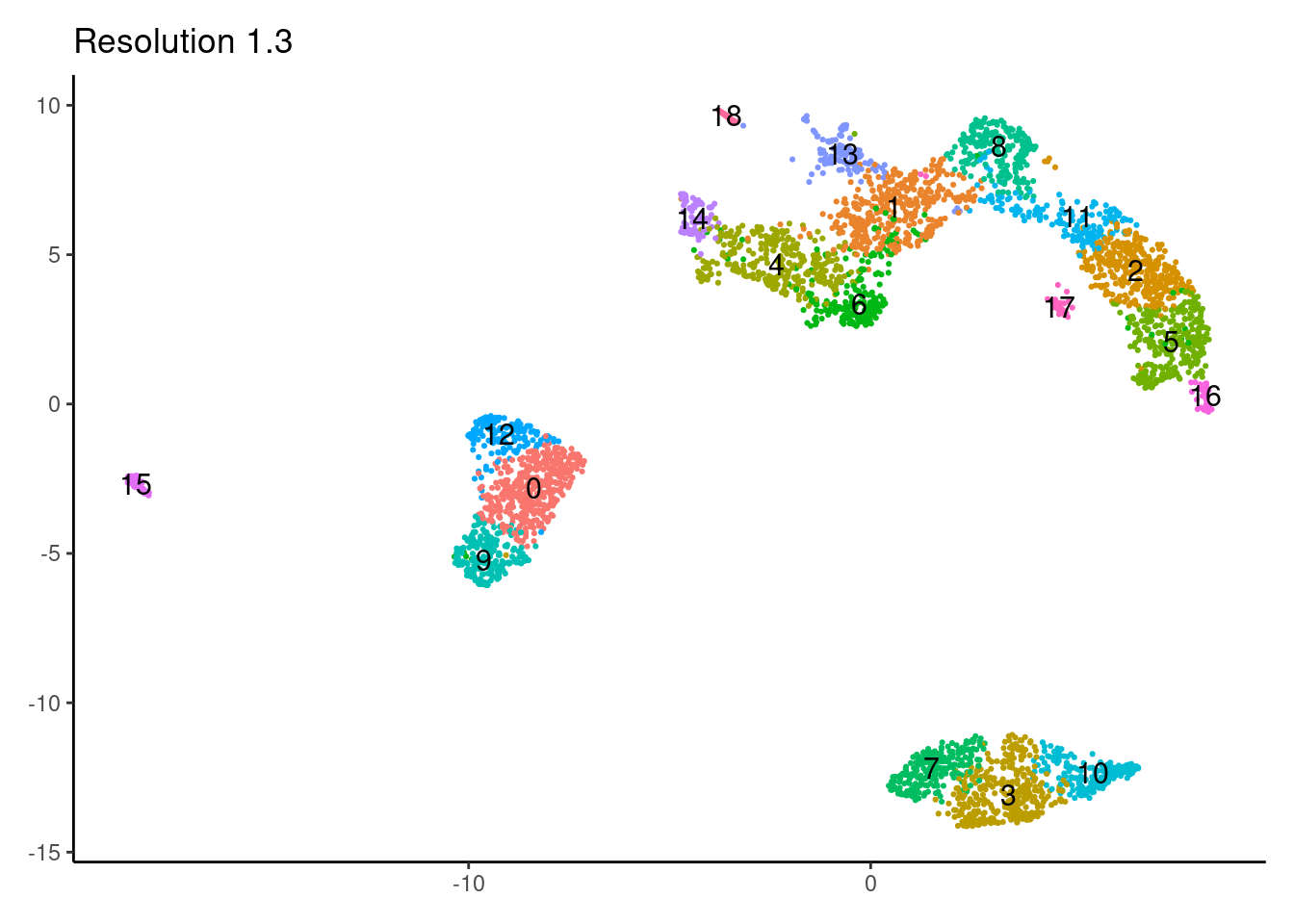
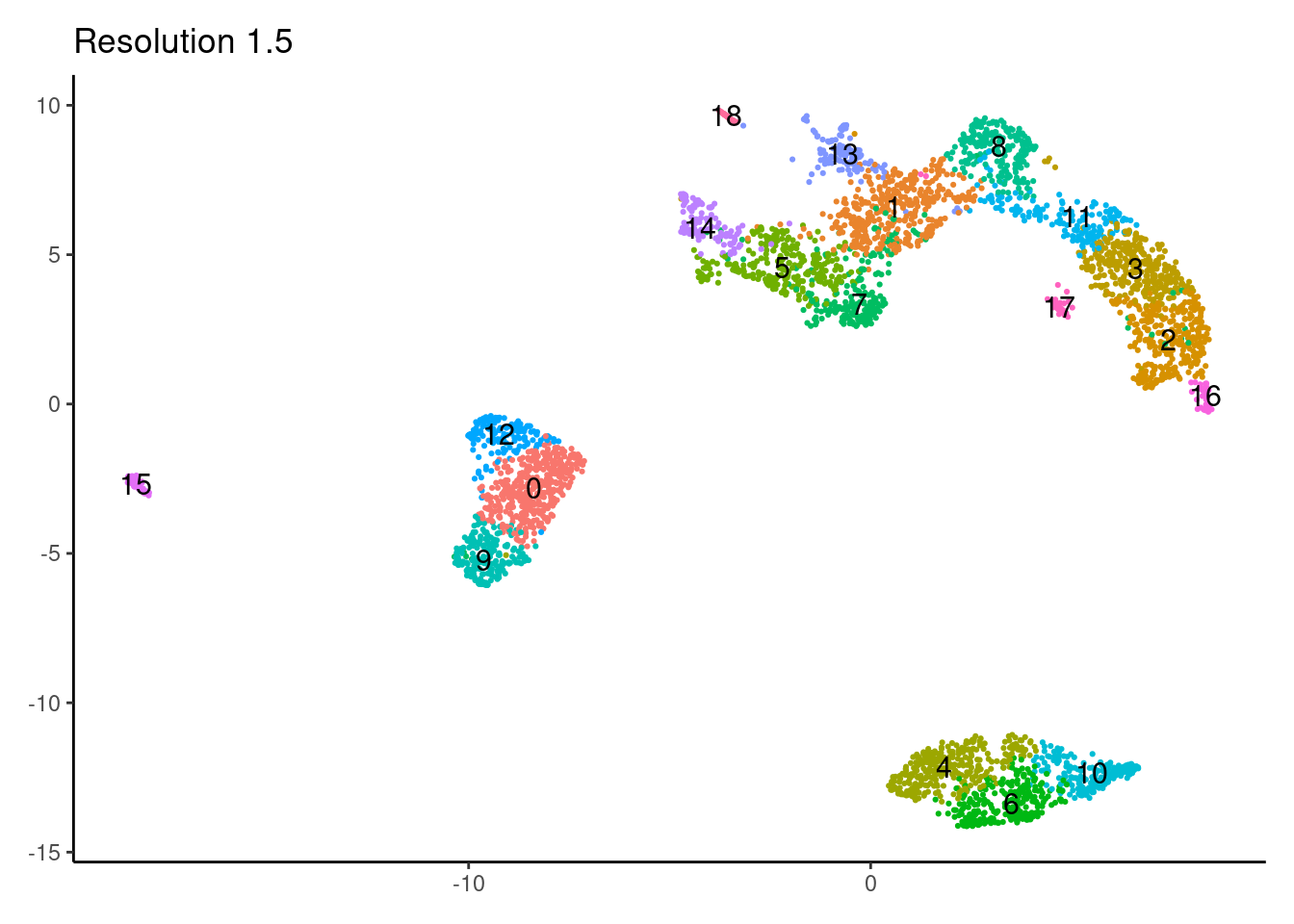
1.5
Clustree Resolutions
clustree(myeloids, prefix = 'RNA_snn_res.') + guides(size = 'none', shape = 'none', edge_colour = FALSE, edge_alpha = FALSE) + theme(legend.position = 'right')## Warning: `guides(<scale> = FALSE)` is deprecated. Please use `guides(<scale> = "none")` instead.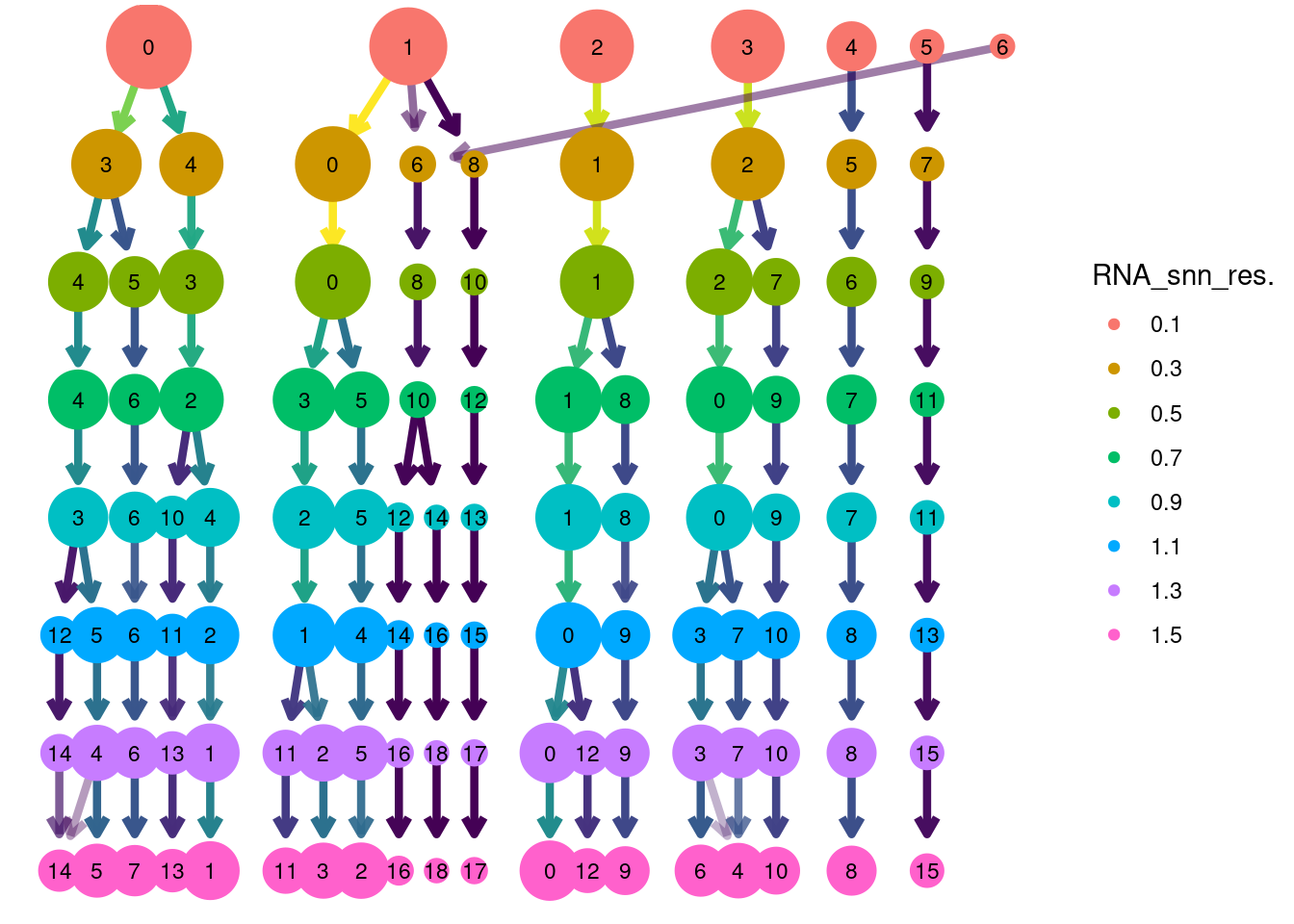
Top2 markers
setwd('~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/SUBSETS/')
for(i in c('0.1','0.3','0.5','0.7','0.9','1.1','1.3','1.5')){
marker_genes <- read_delim(
paste0("Markers/Myeloids/Markers_Myeloids__markers_resolution_",i,".csv"),
delim = ";", escape_double = FALSE, locale = locale(decimal_mark = ",",
grouping_mark = "."), trim_ws = TRUE)
top2 <- marker_genes %>%
dplyr::group_by(cluster)%>%
dplyr::slice(1:2)
top2_g <- unique(top2$gene)
j <- DotPlot(myeloids, features = top2_g, group.by = paste0('RNA_snn_res.', i)) +
theme_classic() +
theme(axis.text.x = element_text(angle=90),
axis.title = element_blank()) +
NoLegend() +
labs(title = paste0('Resolution ',i))
cat("##### ", i, "\n"); print(j); cat("\n\n")
}## Rows: 4201 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.0.1
## Rows: 4494 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.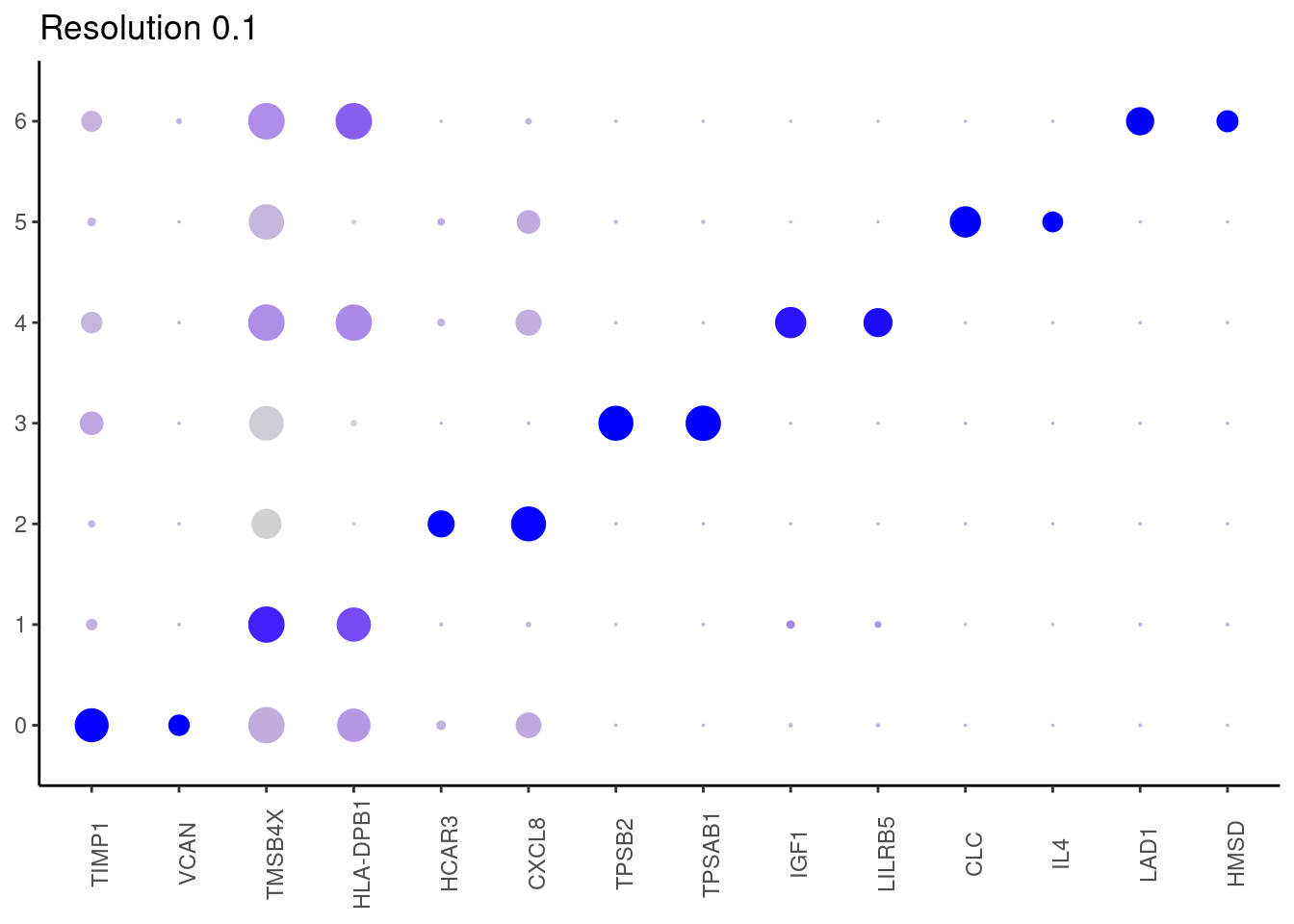 #####
0.3
#####
0.3
## Rows: 5330 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.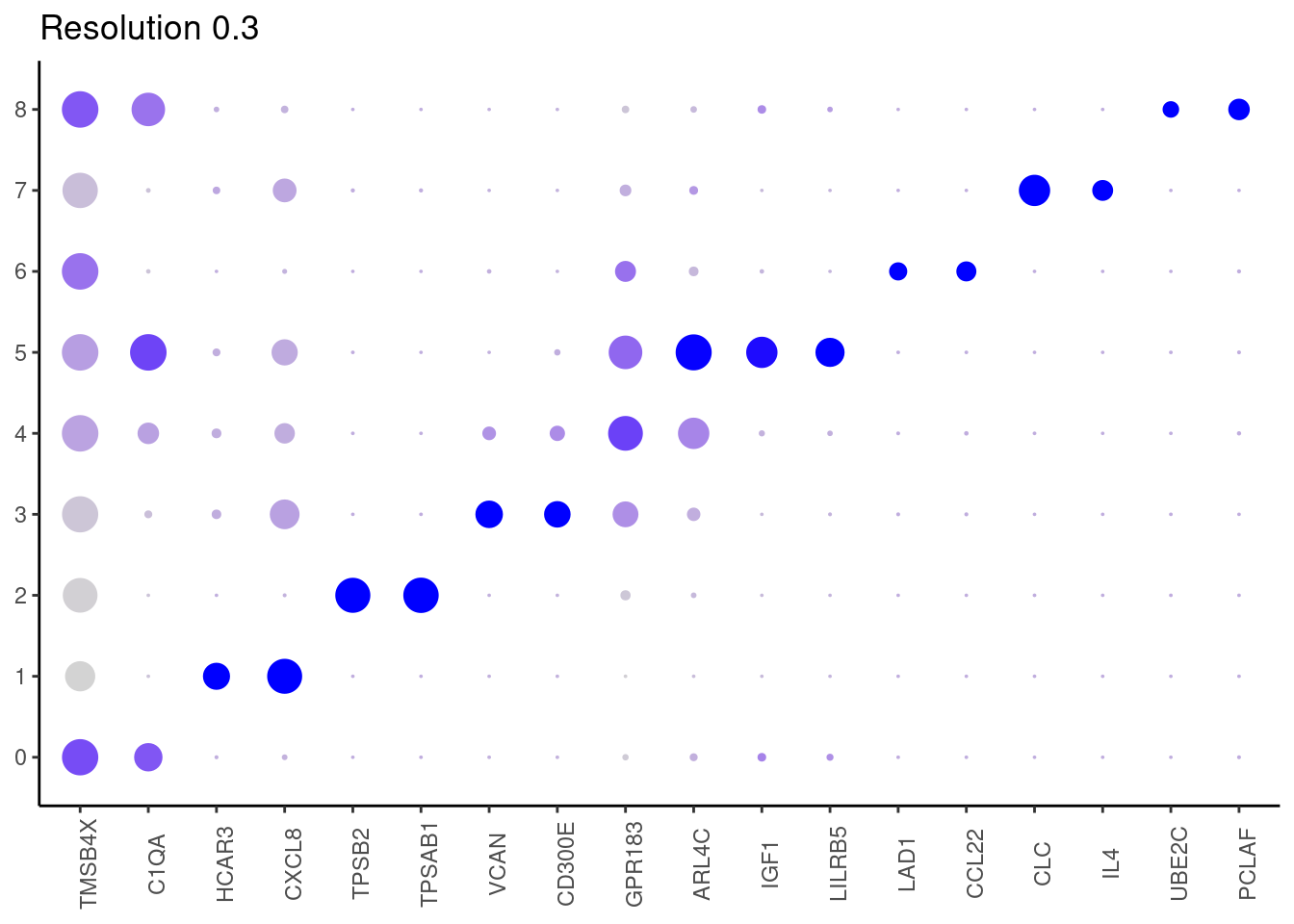 #####
0.5
#####
0.5
## Rows: 5631 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.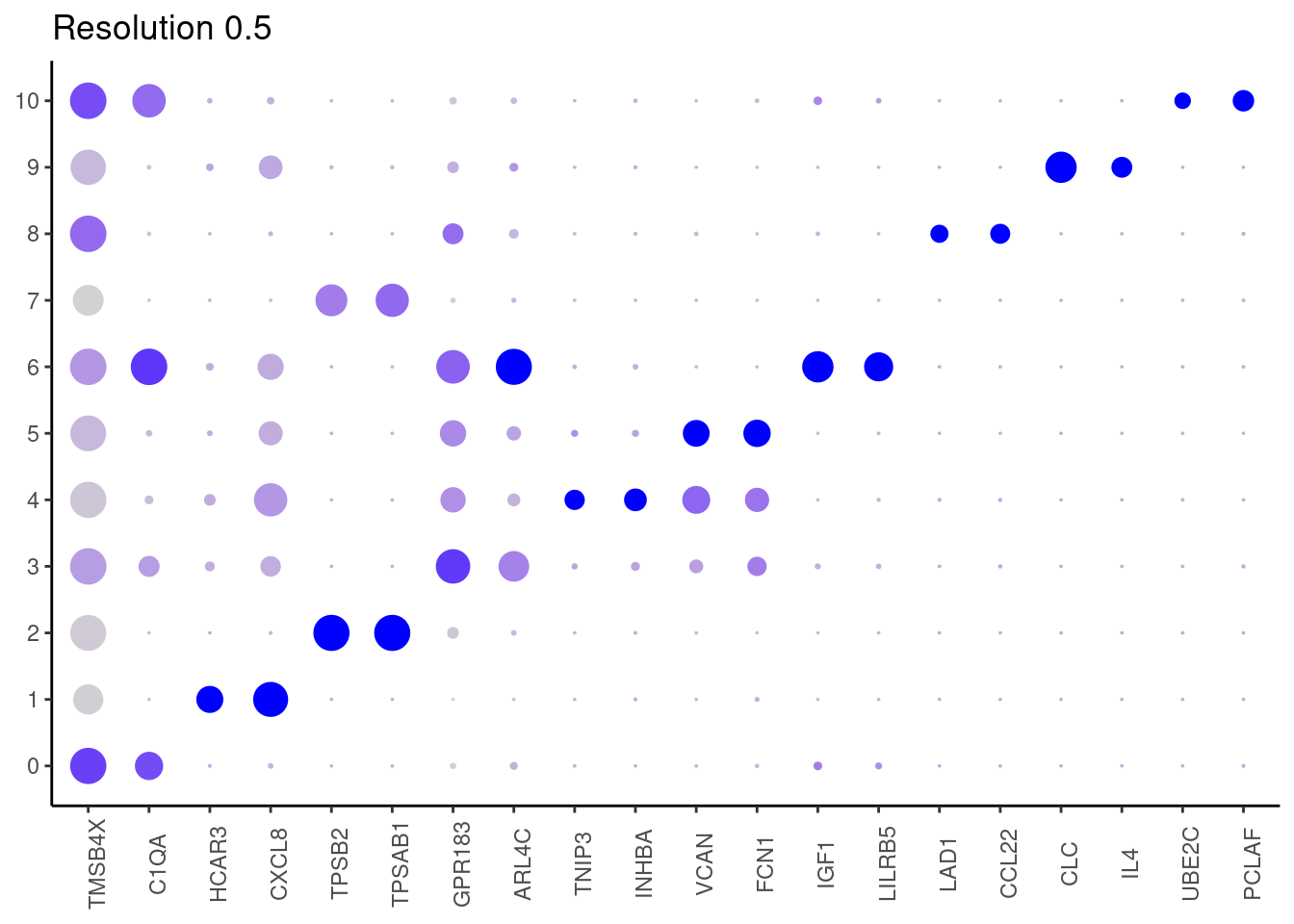 #####
0.7
#####
0.7
## Rows: 7156 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.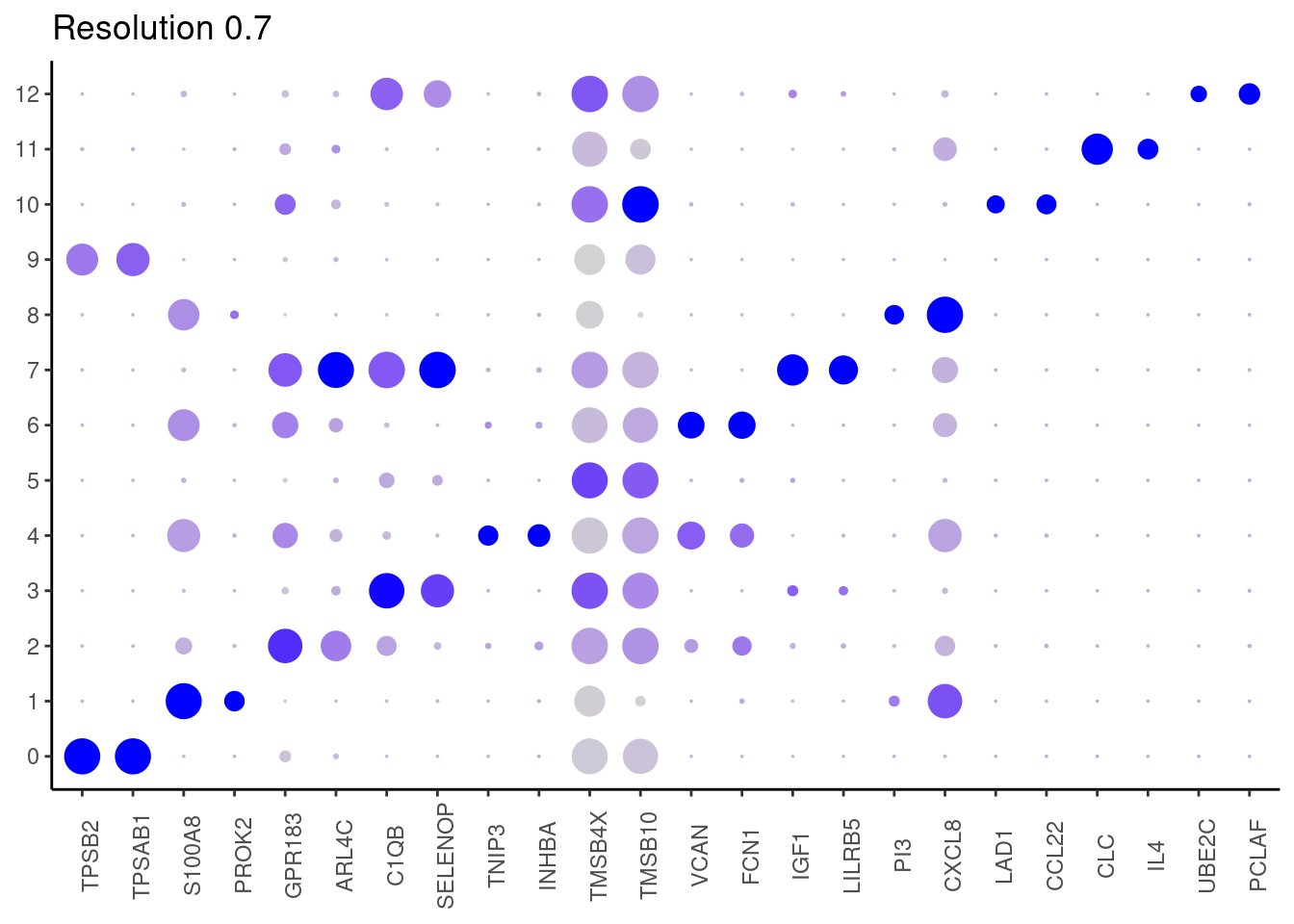 #####
0.9
#####
0.9
## Rows: 8583 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.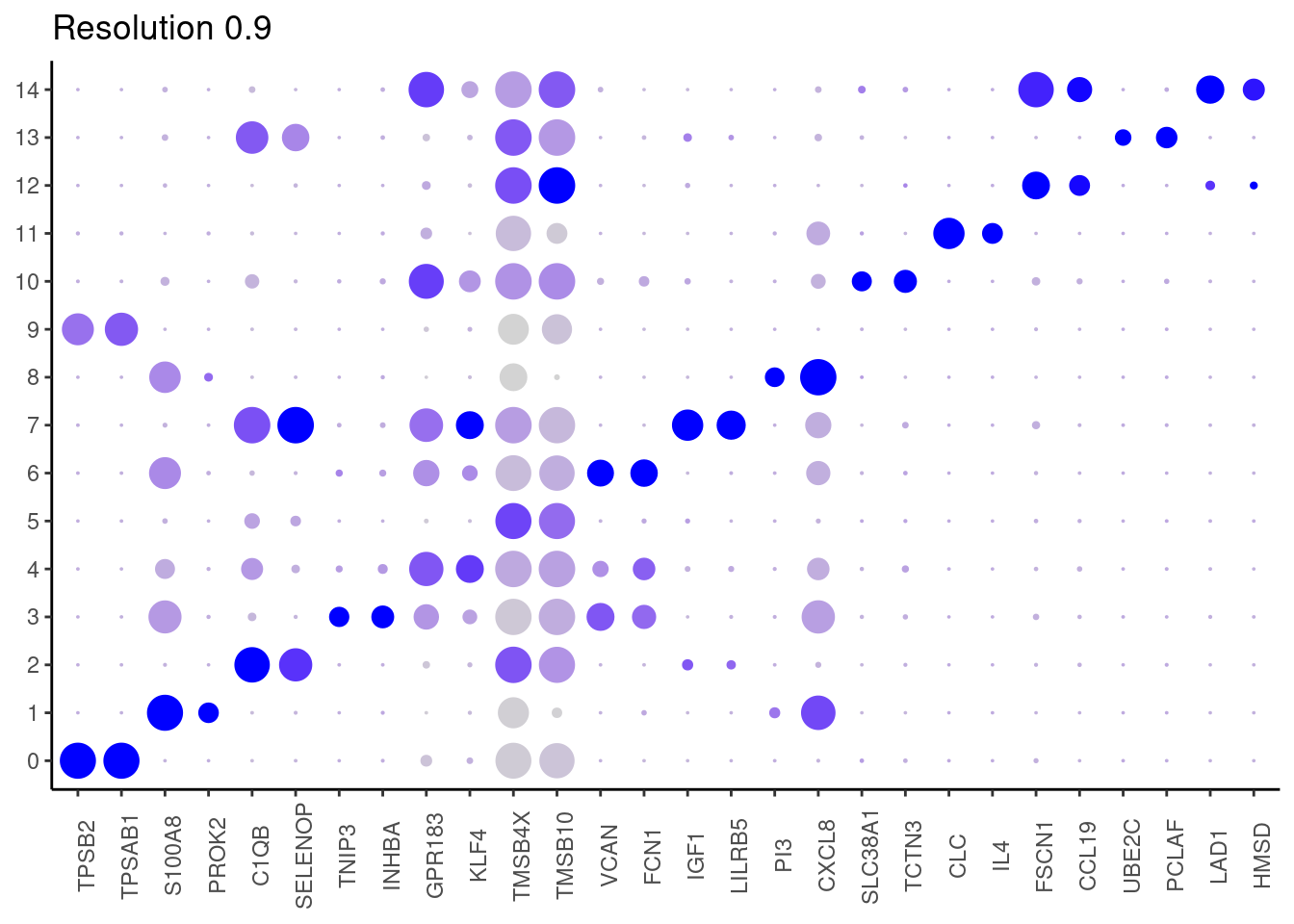 #####
1.1
#####
1.1
## Rows: 9274 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.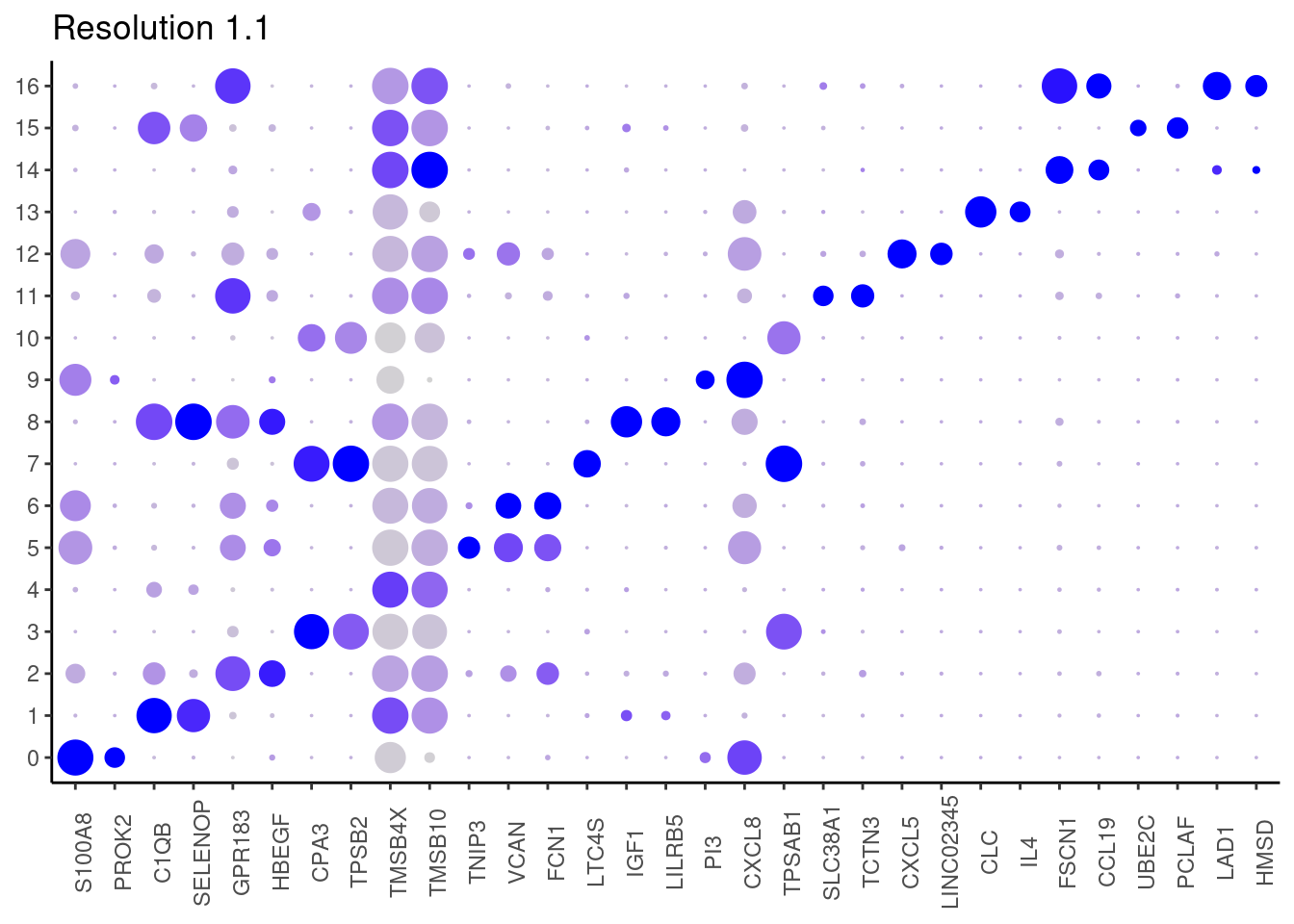 #####
1.3
#####
1.3
## Rows: 9075 Columns: 7
## ── Column specification ──────────────────────────────────────────────────────────────────────────────────────────────
## Delimiter: ";"
## chr (1): gene
## dbl (6): p_val, avg_log2FC, pct.1, pct.2, p_val_adj, cluster
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
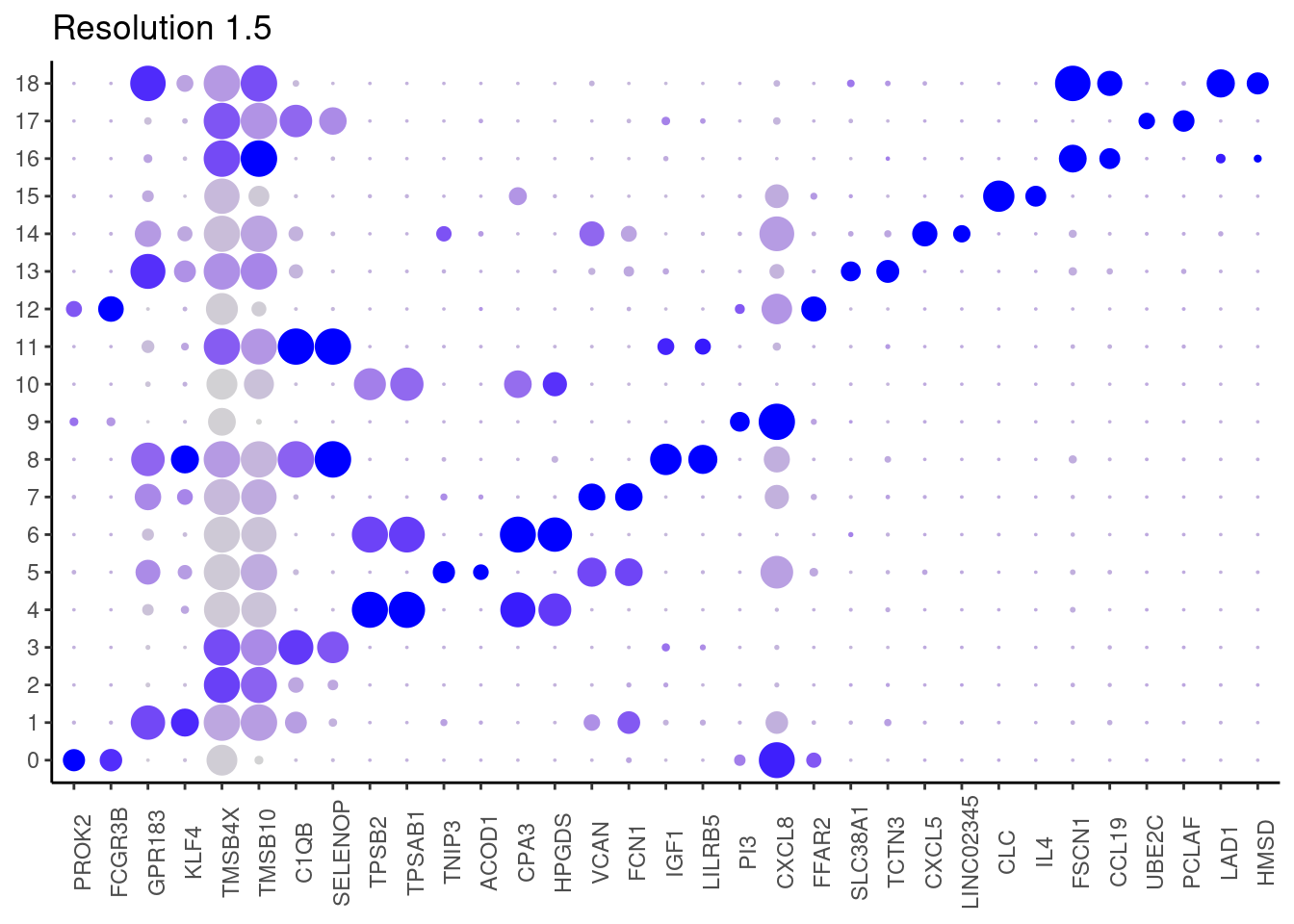
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.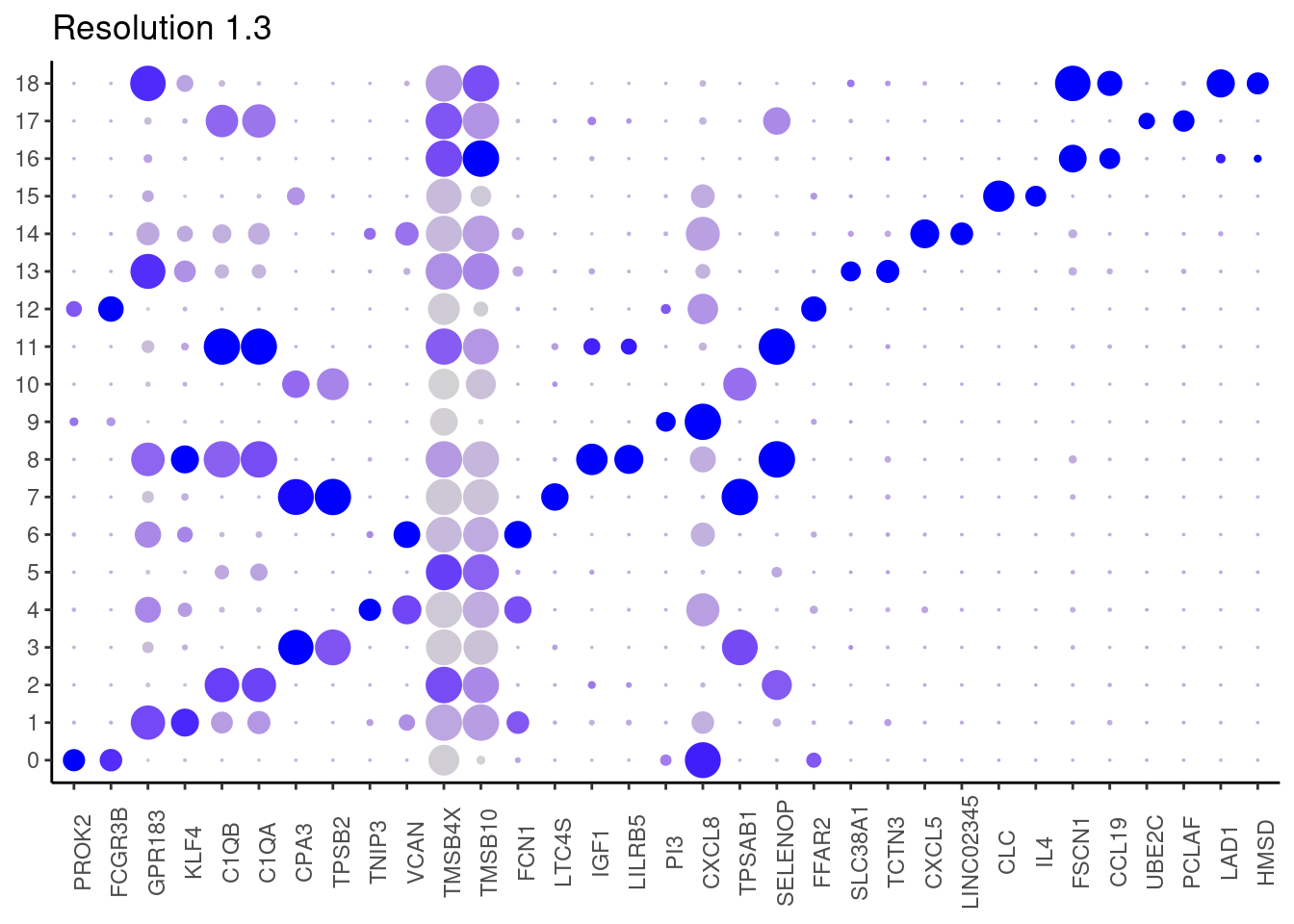 #####
1.5
#####
1.5
Annotation
myeloids$annotation_refined <- plyr::mapvalues(x = myeloids$RNA_snn_res.1.5,
from = 0:18,
to = c("Neutrophil 1",
"IDA macrophage",
"M0 Ribhi",
"M0",
"Mast 1",
"M1 ACOD1",
"Mast 2",
"Inflammatory monocytes",
"M2",
"Neutrophil 2",
"Mast Ribhi",
"M2.2",
"Neutrophil 3",
"DCs CD1c",
"M1 CXCL5",
"Eosinophils",
"DCs CCL22 Ribhi",
"Cycling myeloid",
"DCs CCL22"))
myeloids$annotation_intermediate <- plyr::mapvalues(x = myeloids$RNA_snn_res.1.5,
from = 0:18,
to = c("Neutrophil",
"IDA macrophage",
"M0",
"M0",
"Mast",
"M1",
"Mast",
"Inflammatory monocytes",
"M2",
"Neutrophil",
"Mast",
"M2",
"Neutrophil",
"DCs",
"M1",
"Eosinophils",
"DCs",
"Cycling myeloid",
"DCs"))
a <- DimPlot(myeloids, group.by = 'annotation_intermediate', label = T, repel=T, cols = cols_myeloids_intermediate) + NoLegend()
b <- DimPlot(myeloids, group.by = 'annotation_refined', label = T, repel=T, cols = cols_myeloids) + NoLegend()
a+b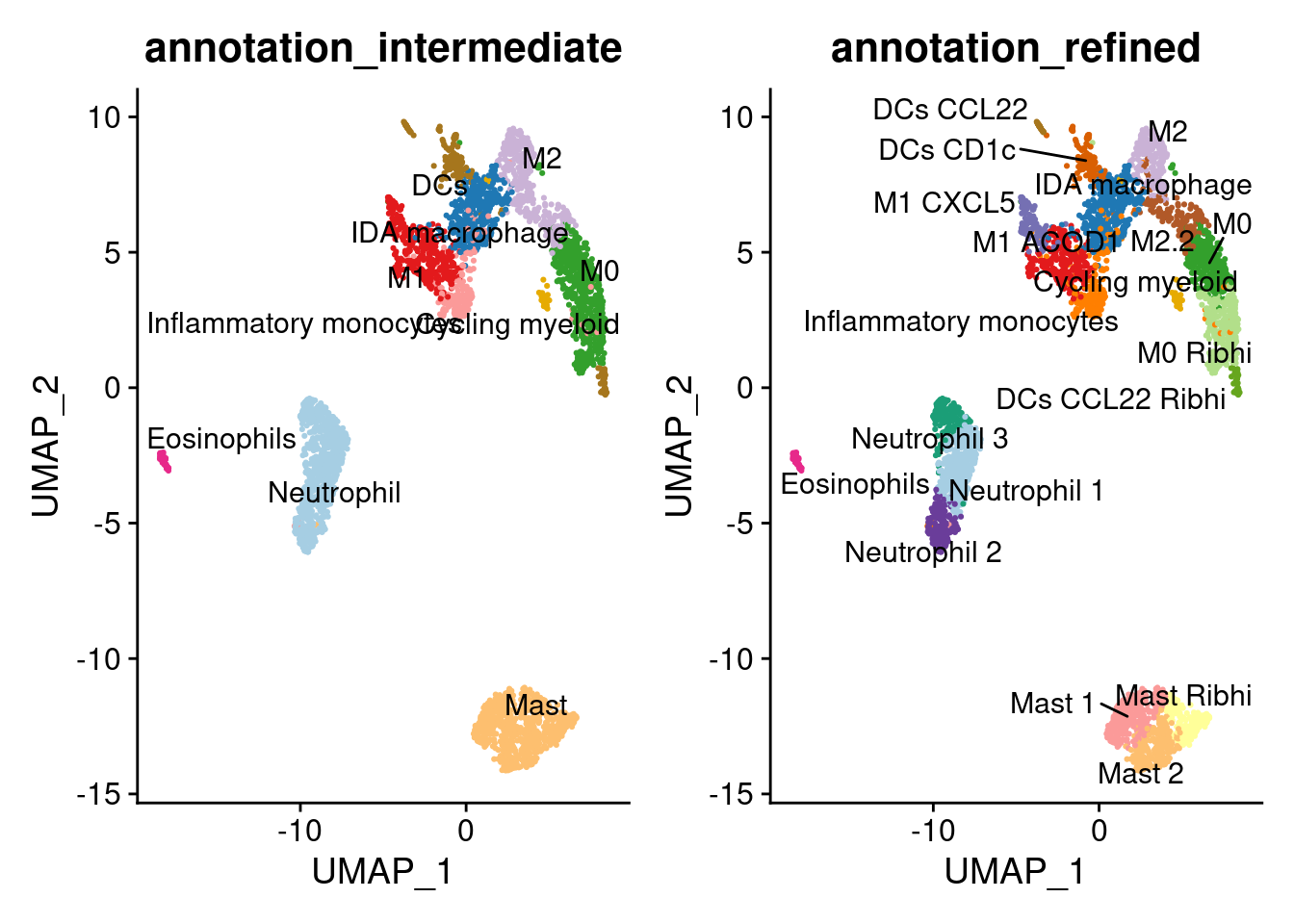
Save file
saveRDS(myeloids, file = '~/000_GitHub/ibd-bcn_single_cell/Analysis of our data/02_Samples_Together/SUBSETS/ON_THEIR_OWN/myeloids_annotated.RDS')
head(myeloids@meta.data)## orig.ident nCount_RNA nFeature_RNA sample doublet Health sample_name Health_2 percent.mt
## SC_002_AACTGGTGTCTCAACA-1 SC 1633 705 SC_002 singlet HC HC 1 HC 2.434571
## SC_002_AAGACCTTCAGCGATT-1 SC 2460 991 SC_002 singlet HC HC 1 HC 6.871463
## SC_002_ACGGGTCCAGCATGAG-1 SC 1154 532 SC_002 singlet HC HC 1 HC 2.842377
## SC_002_ACTGAACGTCCTCTTG-1 SC 3567 1231 SC_002 singlet HC HC 1 HC 2.682313
## SC_002_AGACGTTGTCTGGAGA-1 SC 1220 558 SC_002 singlet HC HC 1 HC 5.479452
## SC_002_AGATCTGTCCAAACTG-1 SC 5136 1496 SC_002 singlet HC HC 1 HC 4.721362
## RNA_snn_res.0.1 seurat_clusters RNA_snn_res.0.3 RNA_snn_res.0.5 RNA_snn_res.0.7
## SC_002_AACTGGTGTCTCAACA-1 3 4 2 2 0
## SC_002_AAGACCTTCAGCGATT-1 1 11 0 0 3
## SC_002_ACGGGTCCAGCATGAG-1 3 4 2 2 0
## SC_002_ACTGAACGTCCTCTTG-1 3 4 2 2 0
## SC_002_AGACGTTGTCTGGAGA-1 1 11 0 0 3
## SC_002_AGATCTGTCCAAACTG-1 4 8 5 6 7
## RNA_snn_res.0.9 RNA_snn_res.1.1 RNA_snn_res.1.3 RNA_snn_res.1.5 annotation_refined
## SC_002_AACTGGTGTCTCAACA-1 0 7 7 4 Mast 1
## SC_002_AAGACCTTCAGCGATT-1 2 1 11 11 M2.2
## SC_002_ACGGGTCCAGCATGAG-1 0 7 7 4 Mast 1
## SC_002_ACTGAACGTCCTCTTG-1 0 7 7 4 Mast 1
## SC_002_AGACGTTGTCTGGAGA-1 2 1 11 11 M2.2
## SC_002_AGATCTGTCCAAACTG-1 7 8 8 8 M2
## annotation_intermediate
## SC_002_AACTGGTGTCTCAACA-1 Mast
## SC_002_AAGACCTTCAGCGATT-1 M2
## SC_002_ACGGGTCCAGCATGAG-1 Mast
## SC_002_ACTGAACGTCCTCTTG-1 Mast
## SC_002_AGACGTTGTCTGGAGA-1 M2
## SC_002_AGATCTGTCCAAACTG-1 M2sessionInfo()
sessionInfo()## R version 4.1.2 (2021-11-01)
## Platform: x86_64-pc-linux-gnu (64-bit)
## Running under: Ubuntu 20.04.5 LTS
##
## Matrix products: default
## BLAS: /usr/lib/x86_64-linux-gnu/blas/libblas.so.3.9.0
## LAPACK: /opt/R/4.1.2/lib/R/lib/libRlapack.so
##
## locale:
## [1] LC_CTYPE=C.UTF-8 LC_NUMERIC=C LC_TIME=C.UTF-8 LC_COLLATE=C.UTF-8
## [5] LC_MONETARY=C.UTF-8 LC_MESSAGES=C LC_PAPER=C.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C LC_MEASUREMENT=C.UTF-8 LC_IDENTIFICATION=C
##
## attached base packages:
## [1] stats4 stats graphics grDevices utils datasets methods base
##
## other attached packages:
## [1] Matrix_1.4-0 nnet_7.3-17 matchSCore2_0.1.0 harmony_0.1.0
## [5] Rcpp_1.0.9 rmarkdown_2.18 pandoc_0.1.0 readxl_1.3.1
## [9] magick_2.7.3 data.table_1.14.2 BiocParallel_1.28.3 RColorBrewer_1.1-3
## [13] ggrepel_0.9.1 ggrastr_1.0.1 usethis_2.1.5 clustree_0.4.4
## [17] ggraph_2.0.5 readr_2.1.2 dplyr_1.0.10 cowplot_1.1.1
## [21] reshape_0.8.8 formulaic_0.0.8 patchwork_1.1.2 MASS_7.3-55
## [25] viridis_0.6.2 viridisLite_0.4.1 scDblFinder_1.8.0 scran_1.22.1
## [29] scater_1.22.0 scuttle_1.4.0 celda_1.10.0 beepr_1.3
## [33] DropletUtils_1.14.2 SingleCellExperiment_1.16.0 SummarizedExperiment_1.24.0 Biobase_2.54.0
## [37] GenomicRanges_1.46.1 GenomeInfoDb_1.30.1 IRanges_2.28.0 S4Vectors_0.32.4
## [41] BiocGenerics_0.40.0 MatrixGenerics_1.6.0 matrixStats_0.62.0 ggplot2_3.3.6
## [45] plyr_1.8.7 sp_1.5-0 SeuratObject_4.1.1 Seurat_4.1.0.9007
##
## loaded via a namespace (and not attached):
## [1] rsvd_1.0.5 ica_1.0-3 corpcor_1.6.10 assertive.properties_0.0-4
## [5] foreach_1.5.2 lmtest_0.9-40 rprojroot_2.0.3 crayon_1.5.2
## [9] spatstat.core_2.4-4 rhdf5filters_1.6.0 backports_1.4.1 nlme_3.1-155
## [13] rlang_1.0.6 XVector_0.34.0 ROCR_1.0-11 irlba_2.3.5
## [17] SparseM_1.81 limma_3.50.3 xgboost_1.5.0.2 rjson_0.2.21
## [21] bit64_4.0.5 glue_1.6.2 sctransform_0.3.4 parallel_4.1.2
## [25] vipor_0.4.5 spatstat.sparse_2.1-1 AnnotationDbi_1.56.2 spatstat.geom_2.4-0
## [29] tidyselect_1.1.2 fitdistrplus_1.1-8 tidyr_1.2.1 assertive.types_0.0-3
## [33] zoo_1.8-10 org.Mm.eg.db_3.14.0 xtable_1.8-4 magrittr_2.0.3
## [37] evaluate_0.18 cli_3.4.0 zlibbioc_1.40.0 rstudioapi_0.13
## [41] miniUI_0.1.1.1 bslib_0.4.1 rpart_4.1.16 RcppEigen_0.3.3.9.2
## [45] shiny_1.7.3 BiocSingular_1.10.0 xfun_0.34 clue_0.3-60
## [49] cluster_2.1.2 tidygraph_1.2.1 KEGGREST_1.34.0 tibble_3.1.8
## [53] listenv_0.8.0 Biostrings_2.62.0 png_0.1-7 future_1.28.0
## [57] withr_2.5.0 bitops_1.0-7 ggforce_0.3.3 cellranger_1.1.0
## [61] assertive.base_0.0-9 dqrng_0.3.0 pillar_1.8.1 GlobalOptions_0.1.2
## [65] cachem_1.0.6 fs_1.5.2 GetoptLong_1.0.5 DelayedMatrixStats_1.16.0
## [69] vctrs_0.4.1 ellipsis_0.3.2 generics_0.1.3 tools_4.1.2
## [73] beeswarm_0.4.0 munsell_0.5.0 tweenr_1.0.2 DelayedArray_0.20.0
## [77] fastmap_1.1.0 compiler_4.1.2 pkgload_1.2.4 abind_1.4-5
## [81] httpuv_1.6.6 plotly_4.10.0 rgeos_0.5-9 GenomeInfoDbData_1.2.7
## [85] gridExtra_2.3 enrichR_3.0 edgeR_3.36.0 lattice_0.20-45
## [89] deldir_1.0-6 utf8_1.2.2 later_1.3.0 jsonlite_1.8.3
## [93] multipanelfigure_2.1.2 scales_1.2.1 graph_1.72.0 ScaledMatrix_1.2.0
## [97] pbapply_1.5-0 sparseMatrixStats_1.6.0 lazyeval_0.2.2 promises_1.2.0.1
## [101] doParallel_1.0.17 R.utils_2.12.0 goftest_1.2-3 checkmate_2.0.0
## [105] spatstat.utils_2.3-1 reticulate_1.26 textshaping_0.3.6 statmod_1.4.36
## [109] Rtsne_0.16 uwot_0.1.14 igraph_1.3.4 HDF5Array_1.22.1
## [113] survival_3.2-13 rsconnect_0.8.25 yaml_2.3.6 systemfonts_1.0.4
## [117] htmltools_0.5.3 memoise_2.0.1 locfit_1.5-9.6 graphlayouts_0.8.0
## [121] digest_0.6.30 assertthat_0.2.1 rappdirs_0.3.3 mime_0.12
## [125] RSQLite_2.2.17 future.apply_1.9.1 blob_1.2.3 R.oo_1.25.0
## [129] ragg_1.2.1 splines_4.1.2 labeling_0.4.2 Rhdf5lib_1.16.0
## [133] RCurl_1.98-1.8 assertive.numbers_0.0-2 hms_1.1.1 rhdf5_2.38.1
## [137] colorspace_2.0-3 ggbeeswarm_0.6.0 shape_1.4.6 assertive.files_0.0-2
## [141] sass_0.4.2 RANN_2.6.1 circlize_0.4.14 audio_0.1-10
## [145] fansi_1.0.3 tzdb_0.2.0 brio_1.1.3 parallelly_1.32.1
## [149] R6_2.5.1 grid_4.1.2 ggridges_0.5.3 lifecycle_1.0.3
## [153] formatR_1.12 bluster_1.4.0 curl_4.3.2 jquerylib_0.1.4
## [157] leiden_0.4.3 testthat_3.1.2 desc_1.4.0 RcppAnnoy_0.0.19
## [161] org.Hs.eg.db_3.14.0 iterators_1.0.14 stringr_1.4.1 topGO_2.46.0
## [165] htmlwidgets_1.5.4 beachmat_2.10.0 polyclip_1.10-0 purrr_0.3.4
## [169] gridGraphics_0.5-1 ComplexHeatmap_2.10.0 mgcv_1.8-38 globals_0.16.1
## [173] spatstat.random_2.2-0 progressr_0.11.0 codetools_0.2-18 GO.db_3.14.0
## [177] metapod_1.2.0 MCMCprecision_0.4.0 R.methodsS3_1.8.2 gtable_0.3.1
## [181] DBI_1.1.3 highr_0.9 tensor_1.5 httr_1.4.4
## [185] KernSmooth_2.23-20 vroom_1.5.7 stringi_1.7.8 reshape2_1.4.4
## [189] farver_2.1.1 combinat_0.0-8 BiocNeighbors_1.12.0 scattermore_0.8
## [193] bit_4.0.4 spatstat.data_2.2-0 pkgconfig_2.0.3 corrplot_0.92
## [197] knitr_1.40